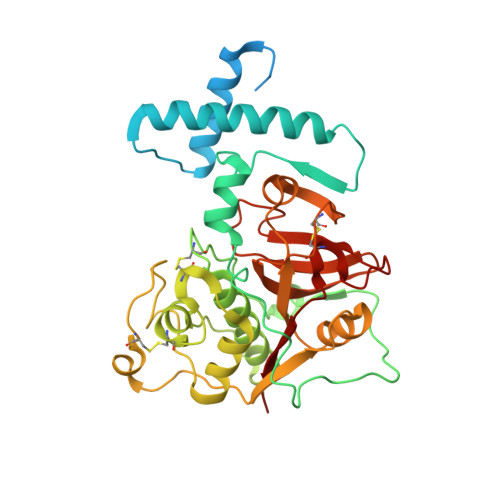
The structure of a thermostable mutant of pro-papain reveals its activation mechanism
Roy, S., Choudhury, D., Aich, P., Dattagupta, J.K., Biswas, S.(2012) Acta Crystallogr D Biol Crystallogr 68: 1591-1603
- PubMed: 23151624 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912038607
- Primary Citation Related Structures:
3TNX - PubMed Abstract:
Papain is the archetype of a broad class of cysteine proteases (clan C1A) that contain a pro-peptide in the zymogen form which is required for correct folding and spatio-temporal regulation of proteolytic activity in the initial stages after expression. This study reports the X-ray structure of the zymogen of a thermostable mutant of papain at 2.6 Å resolution. The overall structure, in particular that of the mature part of the protease, is similar to those of other members of the family. The structure provides an explanation for the molecular basis of the maintenance of latency of the proteolytic activity of the zymogen by its pro-segment at neutral pH. The structural analysis, together with biochemical and biophysical studies, demonstrated that the pro-segment of the zymogen undergoes a rearrangement in the form of a structural loosening at acidic pH which triggers the proteolytic activation cascade. This study further explains the bimolecular stepwise autocatalytic activation mechanism by limited proteolysis of the zymogen of papain at the molecular level. The possible factors responsible for the higher thermal stability of the papain mutant have also been analyzed.
- Crystallography and Molecular Biology Division, Saha Institute of Nuclear Physics, 1/AF Bidhannagar, Kolkata, India.
Organizational Affiliation: