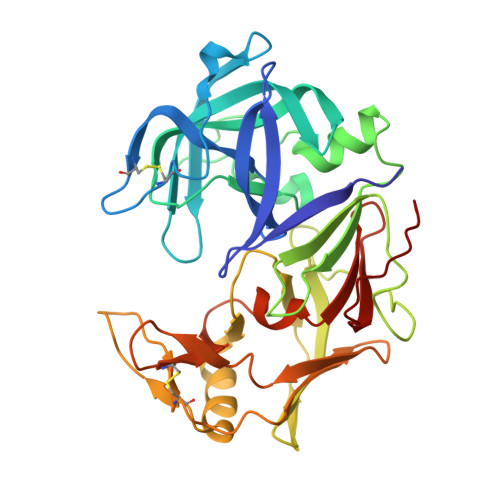
The crystal structure of protease Sapp1p from Candida parapsilosis in complex with the HIV protease inhibitor ritonavir.
Dostal, J., Brynda, J., Hruskova-Heidingsfeldova, O., Pachl, P., Pichova, I., Rezacova, P.(2012) J Enzyme Inhib Med Chem 27: 160-165
- PubMed: 22146051 Search on PubMed
- DOI: https://doi.org/10.3109/14756366.2011.627508
- Primary Citation Related Structures:
3TNE - PubMed Abstract:
Secreted aspartic proteases (Saps) are extracellular proteolytic enzymes that enhance the virulence of Candida pathogens. These enzymes therefore represent possible targets for therapeutic drug design. Saps are inhibited by nanomolar concentrations of the classical inhibitor of aspartic proteases pepstatin A and also by the inhibitors of the HIV protease, but with the K(i) of micromolar values or higher. To contribute to the discussion regarding whether HIV protease inhibitors can act against opportunistic mycoses by the inhibition of Saps, we determined the structure of Sapp1p from Candida parapsilosis in complex with ritonavir (RTV), a clinically used inhibitor of the HIV protease. The crystal structure refined at resolution 2.4 Å proved binding of RTV into the active site of Sapp1p and provided the structural information necessary to evaluate the stability and specificity of the protein-inhibitor interaction.
- Institute of Organic Chemistry and Biochemistry, Academy of Sciences of the Czech Republic , v.v.i., Prague, Czech Republic.
Organizational Affiliation: