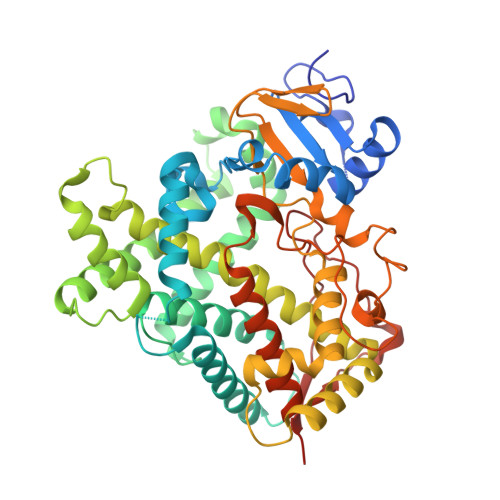
Conformational Adaptation of Human Cytochrome P450 2B6 and Rabbit Cytochrome P450 2B4 Revealed upon Binding Multiple Amlodipine Molecules.
Shah, M.B., Wilderman, P.R., Pascual, J., Zhang, Q., Stout, C.D., Halpert, J.R.(2012) Biochemistry 51: 7225-7238
- PubMed: 22909231 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi300894z
- Primary Citation Related Structures:
3TMZ, 3UA5 - PubMed Abstract:
Structures of human cytochrome P450 2B6 and rabbit cytochrome P450 2B4 in complex with two molecules of the calcium channel blocker amlodipine have been determined by X-ray crystallography. The presence of two drug molecules suggests clear substrate access channels in each P450. According to a previously established nomenclature, amlodipine molecules were trapped in access pathway 2f in P450 2B6 and in pathway 2a or 2f in P450 2B4. These pathways overlap for part of the length and then diverge as they extend toward the protein surface. A previously described solvent channel was also found in each enzyme. The results indicate that key residues located on the surface and at the entrance of the substrate access channels in each of these P450s may play a crucial role in guiding substrate entry. In addition, the region of P450 2B6 and 2B4 involving helices B', F, F', and G' and part of helix G is substantially more open in the amlodipine complexes than in the corresponding 4-(4-chlorophenyl)imidazole complexes. The increased active site volume observed results from the major retraction of helices F, F', and B' and the β4 sheet region located close to the binding cavity to accommodate amlodipine. These structures demonstrate novel insight into distinct conformational states not observed with previous P450 2B structures and provide clear evidence of the substrate access channels in two drug-metabolizing P450s. In addition, the structures exhibit the versatility that can be exploited via in silico studies with other P450 2B6 ligands as large as raloxifene and itraconazole.
- Skaggs School of Pharmacy and Pharmaceutical Sciences, University of California, San Diego, La Jolla, California 92093, United States. m7shah@ucsd.edu
Organizational Affiliation: