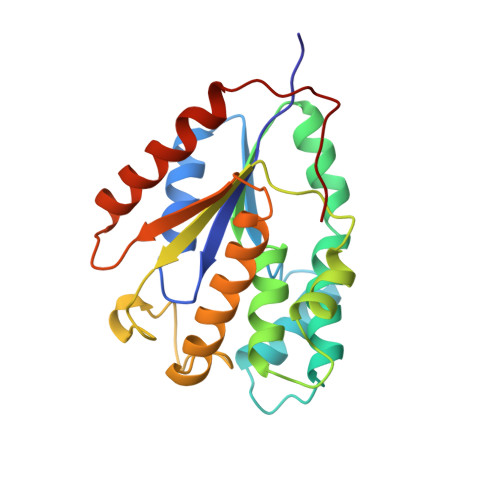
Crystal structure of yeast thymidylate kinase complexed with the bisubstrate inhibitor P1-(5'-adenosyl) P5-(5'-thymidyl) pentaphosphate (TP5A) at 2.0 A resolution: implications for catalysis and AZT activation.
Lavie, A., Konrad, M., Brundiers, R., Goody, R.S., Schlichting, I., Reinstein, J.(1998) Biochemistry 37: 3677-3686
- PubMed: 9521686 Search on PubMed
- DOI: https://doi.org/10.1021/bi9720787
- Primary Citation Related Structures:
3TMK - PubMed Abstract:
The crystal structure of yeast thymidylate kinase (TmpK) complexed with the bisubstrate inhibitor P1-(5'-adenosyl) P5-(5'-thymidyl) pentaphosphate (TP5A) was determined at 2.0 A resolution. In this complex, TmpK adopts a closed conformation with a region (LID) of the protein closing upon the substrate and forming a helix. The interactions of TmpK and TP5A strongly suggest that arginine 15, which is located in the phosphate binding loop (P-loop) sequence, plays a catalytic role by interacting with an oxygen atom of the transferred phosphoryl group. Unlike other nucleoside monophosphate kinases where basic residues from the LID region participate in stabilizing the transition state, TmpK lacks such residues in the LID region. We attribute this function to Arg 15 of the P-loop. TmpK plays an important role in the phosphorylation of the AIDS prodrug AZT. The structures of TmpK with dTMP and with AZT-MP [Lavie, A., et al. (1997) Nat. Struct. Biol. 4, 601-604] implicate the movement of Arg15 in response to AZT-MP binding as an important factor in the 200-fold reduced catalytic rate with AZT-MP. TmpK from Escherichia coli lacks this arginine in its P-loop while having basic residues in the LID region. This suggested that, if such a P-loop movement were to occur in the E. coli TmpK upon AZT-MP binding, it should not have such a detrimental effect on catalysis. This hypothesis was tested, and as postulated, E. coli TmpK phosphorylates AZT-MP only 2.5 times slower than dTMP.
- Department of Physical Biochemistry, Max Planck Institute for Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: