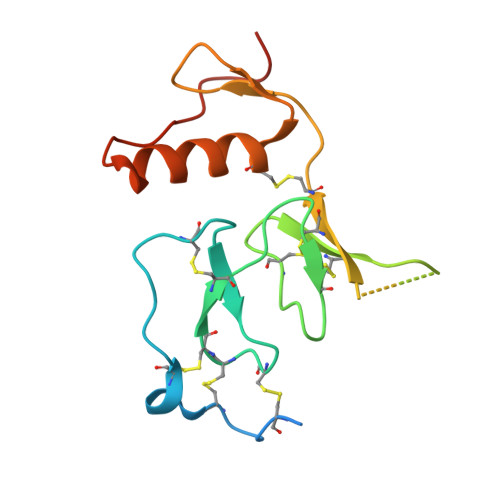
Structural and Functional Analysis of HtrA1 and Its Subdomains.
Eigenbrot, C., Ultsch, M., Lipari, M.T., Moran, P., Lin, S.J., Ganesan, R., Quan, C., Tom, J., Sandoval, W., van Lookeren Campagne, M., Kirchhofer, D.(2012) Structure 20: 1040-1050
- PubMed: 22578544 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.03.021
- Primary Citation Related Structures:
3TJN, 3TJO, 3TJQ - PubMed Abstract:
The homotrimeric human serine protease HtrA1 is homologous to bacterial HtrA proteases regarding the trypsin-like catalytic and PDZ domains but differs by the presence of an N-terminal domain with IGFBP and Kazal homology. The crystal structures and SAXS analysis presented herein reveal the rare tandem of IGFBP- and Kazal-like modules, a protease active site that adopts a competent conformation in the absence of substrate or inhibitor and a model for the intact protein in solution. Highly sensitive enzymatic assays and binding studies demonstrate that the N-terminal tandem has no apparent effect on protease activity, and in accordance with the structure-based predictions, neither the IGFBP- nor Kazal-like module retains the function of their prototype proteins. Our structures of the unliganded HtrA1 active site suggest two-state equilibrium and a "conformational selection" model, in which substrate binds to the active conformer.
- Department of Structural Biology, Genentech, Inc., South San Francisco, CA 94080, USA.
Organizational Affiliation: