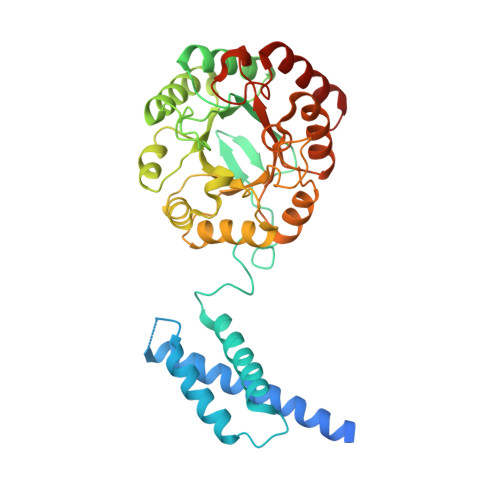
Structural analysis of a 3-deoxy-D-arabino-heptulosonate 7-phosphate synthase with an N-terminal chorismate mutase-like regulatory domain.
Light, S.H., Halavaty, A.S., Minasov, G., Shuvalova, L., Anderson, W.F.(2012) Protein Sci 21: 887-895
- PubMed: 22505283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2075
- Primary Citation Related Structures:
3NVT, 3TFC - PubMed Abstract:
3-Deoxy-D-arabino-heptulosonate 7-phosphate synthase (DAHPS) catalyzes the first step in the biosynthesis of a number of aromatic metabolites. Likely because this reaction is situated at a pivotal biosynthetic gateway, several DAHPS classes distinguished by distinct mechanisms of allosteric regulation have independently evolved. One class of DAHPSs contains a regulatory domain with sequence homology to chorismate mutase-an enzyme further downstream of DAHPS that catalyzes the first committed step in tyrosine/phenylalanine biosynthesis-and is inhibited by chorismate mutase substrate (chorismate) and product (prephenate). Described in this work, structures of the Listeria monocytogenes chorismate/prephenate regulated DAHPS in complex with Mn(2+) and Mn(2+) + phosphoenolpyruvate reveal an unusual quaternary architecture: DAHPS domains assemble as a tetramer, from either side of which chorismate mutase-like (CML) regulatory domains asymmetrically emerge to form a pair of dimers. This domain organization suggests that chorismate/prephenate binding promotes a stable interaction between the discrete regulatory and catalytic domains and supports a mechanism of allosteric inhibition similar to tyrosine/phenylalanine control of a related DAHPS class. We argue that the structural similarity of chorismate mutase enzyme and CML regulatory domain provides a unique opportunity for the design of a multitarget antibacterial.
- Center for Structural Genomics of Infectious Diseases, Department of Molecular Pharmacology and Biological Chemistry, Feinberg School of Medicine, Northwestern University, Chicago, Illinois 60611, USA.
Organizational Affiliation: