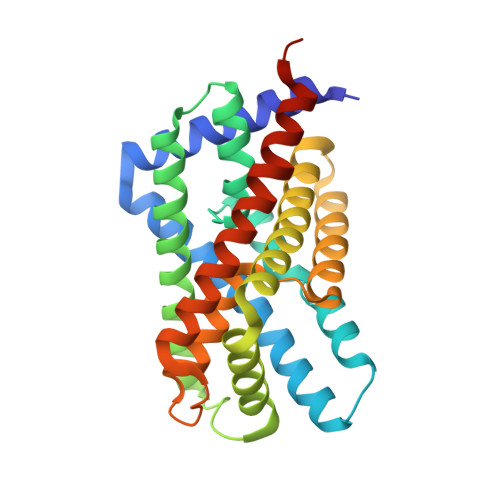
Identification and characterization of a bacterial hydrosulphide ion channel.
Czyzewski, B.K., Wang, D.N.(2012) Nature 483: 494-497
- PubMed: 22407320 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature10881
- Primary Citation Related Structures:
3TDO, 3TDP, 3TDR, 3TDS, 3TDX, 3TE0, 3TE1, 3TE2 - PubMed Abstract:
The hydrosulphide ion (HS(-)) and its undissociated form, hydrogen sulphide (H(2)S), which are believed to have been critical to the origin of life on Earth, remain important in physiology and cellular signalling. As a major metabolite in anaerobic bacterial growth, hydrogen sulphide is a product of both assimilatory and dissimilatory sulphate reduction. These pathways can reduce various oxidized sulphur compounds including sulphate, sulphite and thiosulphate. The dissimilatory sulphate reduction pathway uses this molecule as the terminal electron acceptor for anaerobic respiration, in which process it produces excess amounts of H(2)S (ref. 4). The reduction of sulphite is a key intermediate step in all sulphate reduction pathways. In Clostridium and Salmonella, an inducible sulphite reductase is directly linked to the regeneration of NAD(+), which has been suggested to have a role in energy production and growth, as well as in the detoxification of sulphite. Above a certain concentration threshold, both H(2)S and HS(-) inhibit cell growth by binding the metal centres of enzymes and cytochrome oxidase, necessitating a release mechanism for the export of this toxic metabolite from the cell. Here we report the identification of a hydrosulphide ion channel in the pathogen Clostridium difficile through a combination of genetic, biochemical and functional approaches. The HS(-) channel is a member of the formate/nitrite transport family, in which about 50 hydrosulphide ion channels form a third subfamily alongside those for formate (FocA) and for nitrite (NirC). The hydrosulphide ion channel is permeable to formate and nitrite as well as to HS(-) ions. Such polyspecificity can be explained by the conserved ion selectivity filter observed in the channel's crystal structure. The channel has a low open probability and is tightly regulated, to avoid decoupling of the membrane proton gradient.
- The Helen L. and Martin S. Kimmel Center for Biology and Medicine at the Skirball Institute of Biomolecular Medicine, New York University School of Medicine, 540 First Avenue, New York, New York 10016, USA.
Organizational Affiliation: