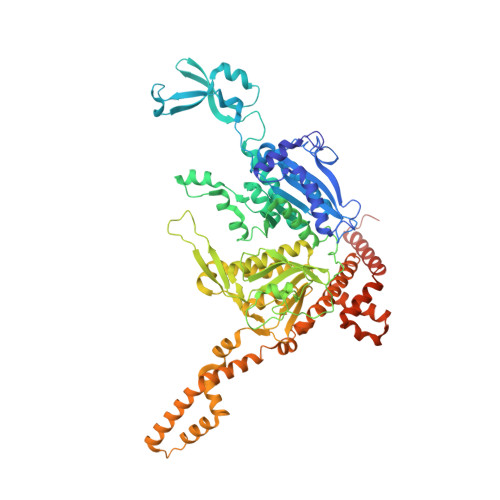
Design, synthesis, and structure-activity relationships of spirolactones bearing 2-ureidobenzothiophene as acetyl-CoA carboxylases inhibitors.
Yamashita, T., Kamata, M., Endo, S., Yamamoto, M., Kakegawa, K., Watanabe, H., Miwa, K., Yamano, T., Funata, M., Sakamoto, J., Tani, A., Mol, C.D., Zou, H., Dougan, D.R., Sang, B., Snell, G., Fukatsu, K.(2011) Bioorg Med Chem Lett 21: 6314-6318
- PubMed: 21944854 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.08.117
- Primary Citation Related Structures:
3TDC - PubMed Abstract:
The co-crystal structure of the human acetyl-coenzyme A 2 (ACC2) carboxyl transferase domain and the reported compound CP-640186 (1b) suggested that two carbonyl groups are essential for potent ACC2 inhibition. By focusing on enhancing the interactions between the two carbonyl groups and the amino acid residues Gly(2162) and Glu(2230), we used ligand- and structure-based drug design to discover spirolactones bearing a 2-ureidobenzothiophene moiety.
- Pharmaceutical Research Division, Takeda Pharmaceutical Company, Ltd, 2-17-85, Jusohonmachi, Yodogawa-ku, Osaka 532-8686, Japan. Yamashita_Tohru@takeda.co.jp
Organizational Affiliation: