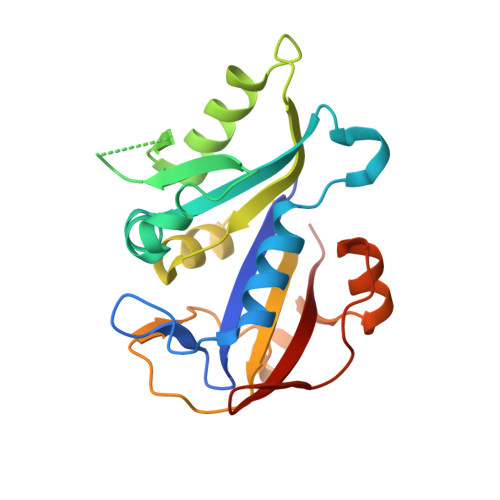
Structural analysis of Pneumocystis cariniidihydrofolate reductase complexed with NADPH and 2,4-diamino-6-[2-(5-carboxypent-1-yn-1-yl)-5-methoxybenzyl]-5-methylpyrido[2,3-d]pyrimidine.
Cody, V., Pace, J., Stewart, E.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 418-423
- PubMed: 22505410 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112008688
- Primary Citation Related Structures:
3TD8 - PubMed Abstract:
Structural data are reported for 2,4-diamino-6-[2-(5-carboxypent-1-yn-1-yl)-5-methoxybenzyl]-5-methylpyrido[2,3-d]pyrimidine (PY1014) complexed with Pneumocystis carinii dihydrofolate reductase (pcDHFR) refined to 1.8 Å resolution. These data reveal that the carboxylate of the ω-carboxyalkynyl side chain of PY1014, the most pcDHFR-selective analog in this series, forms ionic interactions with the conserved Arg75 in the substrate-binding pocket of pcDHFR. The reversal of the 2',5'-substitution pattern of this analog compared with the highly selective diaminopyrimidine analog PY1011 (i.e. the 5'-pentynylcarboxy-5'-methoxy pattern of PY1014 versus the 3',4'-dimethoxy-5'-pentynylcarboxy pattern of PY1011) is necessary to achieve optimal interaction with Arg75 as observed in other structures. The larger diaminopyrido[2,3-d]pyrimidine ring of PY1014 places the 5'-methoxy group closer to Leu25 and Ser64 than does the diaminopyrimidine ring of PY1011. The 5'-methoxy O atom forms a hydrogen bond to the amide of Leu25 (O···N, 2.7 Å) and the 5'-methoxy methyl group makes a hydrophobic contact of 3.1 Å with C(β) of Ser64. Although the IC(50) values of PY1014 and PY1011 are similar, inhibition data show that the selectivity of PY1011 for pcDHFR is significantly greater. The greater selectivity for pcDHFR compared with mammalian DHFR of these inhibitors is also influenced by the enhanced hydrophobic interactions of the side-chain methylene atoms with Phe69 of pcDHFR compared with Asn64 of mammalian DHFR.
- Structural Biology Department, Hauptman-Woodward Medical Research Institute, 700 Ellicott Street, Buffalo, NY 14203, USA. cody@hwi.buffalo.edu
Organizational Affiliation: