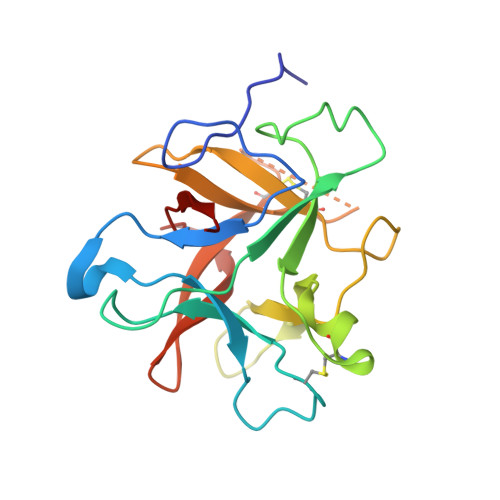
Structure of a post-translationally processed heterodimeric double-headed Kunitz-type serine protease inhibitor from potato.
Meulenbroek, E.M., Thomassen, E.A., Pouvreau, L., Abrahams, J.P., Gruppen, H., Pannu, N.S.(2012) Acta Crystallogr D Biol Crystallogr 68: 794-799
- PubMed: 22751664 Search on PubMed
- DOI: https://doi.org/10.1107/S090744491201222X
- Primary Citation Related Structures:
3TC2 - PubMed Abstract:
Potato serine protease inhibitor (PSPI) constitutes about 22% of the total amount of proteins in potato tubers (cv. Elkana), making it the most abundant protease inhibitor in the plant. PSPI is a heterodimeric double-headed Kunitz-type serine protease inhibitor that can tightly and simultaneously bind two serine proteases by mimicking the substrate of the enzyme with its reactive-site loops. Here, the crystal structure of PSPI is reported, representing the first heterodimeric double-headed Kunitz-type serine protease inhibitor structure to be determined. PSPI has a β-trefoil fold and, based on the structure, two reactive-site loops bearing residues Phe75 and Lys95 were identified.
- Biophysical Structural Chemistry, Leiden University, Einsteinweg 55, 2333 CC Leiden, The Netherlands.
Organizational Affiliation: