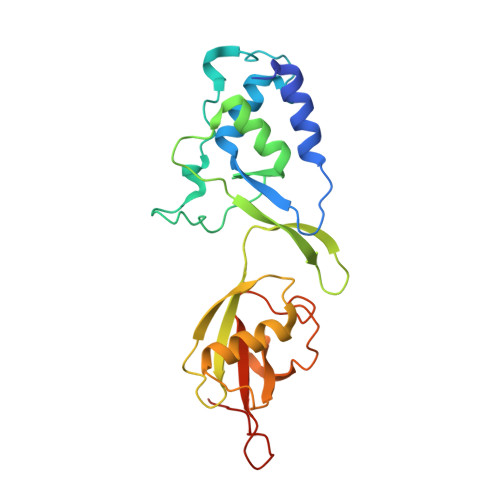
Structure of the USP15 N-Terminal Domains: A beta-Hairpin Mediates Close Association between the DUSP and UBL Domains
Harper, S., Besong, T.M., Emsley, J., Scott, D.J., Dreveny, I.(2011) Biochemistry 50: 7995-8004
- PubMed: 21848306
- DOI: https://doi.org/10.1021/bi200726e
- Primary Citation Related Structures:
3T9L - PubMed Abstract:
Ubiquitin specific protease 15 (USP15) functions in COP9 signalosome mediated regulation of protein degradation and cellular signaling through catalyzing the ubiquitin deconjugation reaction of a discrete number of substrates. It influences the stability of adenomatous polyposis coli, IκBα, caspase-3, and the human papillomavirus type 16 E6. USP15 forms a subfamily with USP4 and USP11 related through a shared presence of N-terminal "domain present in ubiquitin specific proteases" (DUSP) and "ubiquitin-like" (UBL) domains (DU subfamily). Here we report the 1.5 Å resolution crystal structure of the human USP15 N-terminal domains revealing a 80 Å elongated arrangement with the DU domains aligned in tandem. This architecture is generated through formation of a defined interface that is dominated by an intervening β-hairpin structure (DU finger) that engages in an intricate hydrogen-bonding network between the domains. The UBL domain is closely related to ubiquitin among β-grasp folds but is characterized by the presence of longer loop regions and different surface characteristics, indicating that this domain is unlikely to act as ubiquitin mimic. Comparison with the related murine USP4 DUSP-UBL crystal structure reveals that the main DU interdomain contacts are conserved. Analytical ultracentrifugation, small-angle X-ray scattering, and gel filtration experiments revealed that USP15 DU is monomeric in solution. Our data provide a framework to advance study of the structure and function of the DU subfamily.
- Centre for Biomolecular Sciences, University of Nottingham, University Park Campus, Nottingham, NG7 2RD, United Kingdom.
Organizational Affiliation: