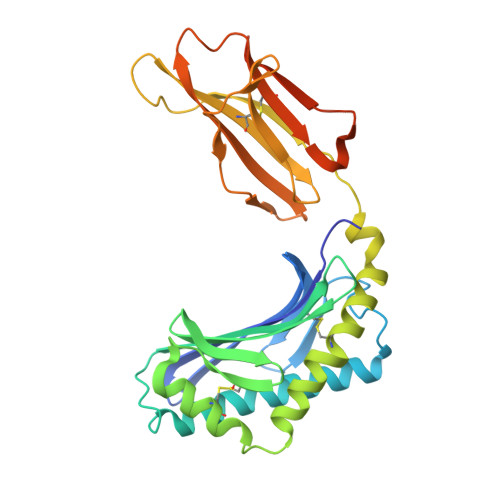
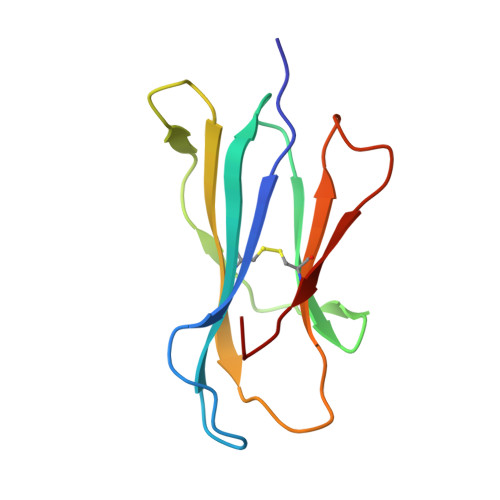
Structural reorganization of the human CD1b Antigen-binding groove for presentation of mycobacterial sulfoglycolipids
Garcia-Alles, L.F., Collmann, A., Versluis, C., Lindner, B., Guiard, J., Maveyraud, L., Huc, E., Im, J.S., Sansano, S., Brando, T., Julien, S., Prandi, J., Gilleron, M., Porcelli, S.A., de la Salle, H., Heck, A.J.R., Mori, L., Puzo, G., Mourey, L., De Libero, G.To be published.