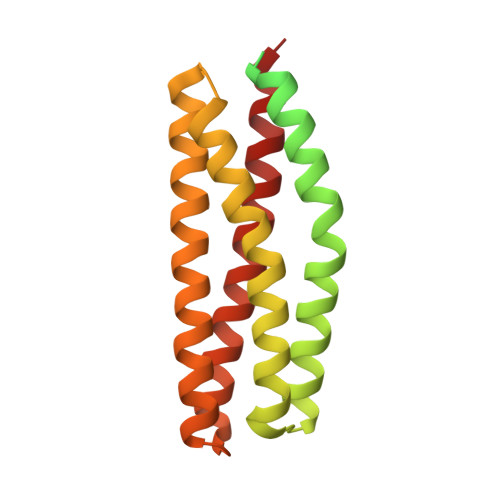
NSP-Cas protein structures reveal a promiscuous interaction module in cell signaling.
Mace, P.D., Wallez, Y., Dobaczewska, M.K., Lee, J.J., Robinson, H., Pasquale, E.B., Riedl, S.J.(2011) Nat Struct Mol Biol 18: 1381-1387
- PubMed: 22081014 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2152
- Primary Citation Related Structures:
3T6A, 3T6G - PubMed Abstract:
Members of the novel SH2-containing protein (NSP) and Crk-associated substrate (Cas) protein families form multidomain signaling platforms that mediate cell migration and invasion through a collection of distinct signaling motifs. Members of each family interact via their respective C-terminal domains, but the mechanism of this association has remained enigmatic. Here we present the crystal structures of the C-terminal domain from the NSP protein BCAR3 and the complex of NSP3 with p130Cas. BCAR3 adopts the Cdc25-homology fold of Ras GTPase exchange factors, but it has a 'closed' conformation incapable of enzymatic activity. The structure of the NSP3-p130Cas complex reveals that this closed conformation is instrumental for interaction of NSP proteins with a focal adhesion-targeting domain present in Cas proteins. This enzyme-to-adaptor conversion enables high-affinity, yet promiscuous, interactions between NSP and Cas proteins and represents an unprecedented mechanistic paradigm linking cellular signaling networks.
- Program of Apoptosis and Cell Death Research, Cancer Center, Sanford-Burnham Medical Research Institute, La Jolla, California, USA.
Organizational Affiliation: