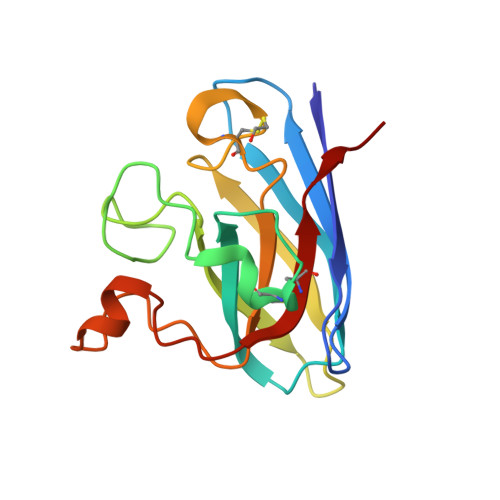
Structural switching of Cu,Zn-superoxide dismutases at loop VI: insights from the crystal structure of 2-mercaptoethanol-modified enzyme
Ihara, K., Fujiwara, N., Yamaguchi, Y., Torigoe, H., Wakatsuki, S., Taniguchi, N., Suzuki, K.(2012) Biosci Rep 32: 539-548
- PubMed: 22804629 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BSR20120029
- Primary Citation Related Structures:
3T5W - PubMed Abstract:
Cu,Zn SOD1 (superoxide dismutase 1) is implicated in FALS (familial amyotrophic lateral sclerosis) through the accumulation of misfolded proteins that are toxic to neuronal cells. Loop VI (residues 102-115) of the protein is at the dimer interface and could play a critical role in stability. The free cysteine residue, Cys111 in the loop, is readily oxidized and alkylated. We have found that modification of this Cys111 with 2-ME (2-mercaptoethanol; 2-ME-SOD1) stabilizes the protein and the mechanism may provide insights into destabilization and the formation of aggregated proteins. Here, we determined the crystal structure of 2-ME-SOD1 and find that the 2-ME moieties in both subunits interact asymmetrically at the dimer interface and that there is an asymmetric configuration of segment Gly108 to Cys111 in loop VI. One loop VI of the dimer forms a 310-helix (Gly108 to His110) within a unique β-bridge stabilized by a hydrogen bond between Ser105-NH and His110-CO, while the other forms a β-turn without the H-bond. The H-bond (H-type) and H-bond free (F-type) configurations are also seen in some wild-type and mutant human SOD1s in the Protein Data Bank suggesting that they are interconvertible and an intrinsic property of SOD1s. The two structures serve as a basis for classification of these proteins and hopefully a guide to their stability and role in pathophysiology.
- Structural Biology Research Center, Institute of Material Structure Science, High Energy Accelerator Research Organization, Tsukuba, Ibaraki, Japan.
Organizational Affiliation: