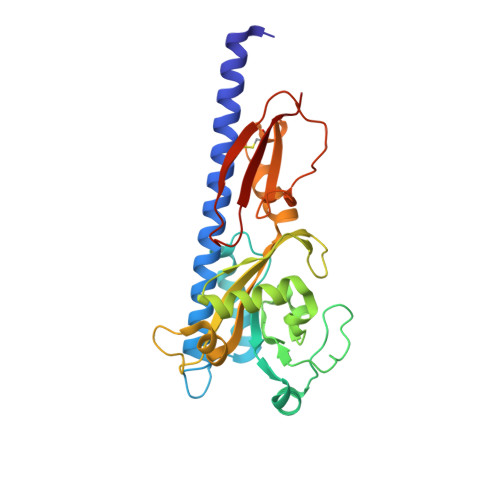
Structural basis for cytokinin recognition by Arabidopsis thaliana histidine kinase 4.
Hothorn, M., Dabi, T., Chory, J.(2011) Nat Chem Biol 7: 766-768
- PubMed: 21964459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.667
- Primary Citation Related Structures:
3T4J, 3T4K, 3T4L, 3T4O, 3T4Q, 3T4S, 3T4T - PubMed Abstract:
Cytokinins are classic hormones that orchestrate plant growth and development and the integrity of stem cell populations. Cytokinin receptors are eukaryotic sensor histidine kinases that are activated by both naturally occurring adenine-type cytokinins and urea-based synthetic compounds. Crystal structures of the Arabidopsis thaliana histidine kinase 4 sensor domain in complex with different cytokinin ligands now rationalize the hormone-binding specificity of the receptor and may spur the design of new cytokinin ligands.
- Plant Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, California, USA.
Organizational Affiliation: