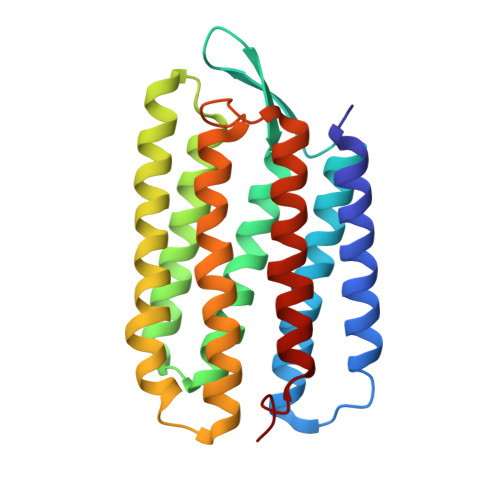
A transporter converted into a sensor, a phototaxis signaling mutant of bacteriorhodopsin at 3.0 angstrom.
Spudich, E.N., Ozorowski, G., Schow, E.V., Tobias, D.J., Spudich, J.L., Luecke, H.(2012) J Mol Biology 415: 455-463
- PubMed: 22123198 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.11.025
- Primary Citation Related Structures:
3T45 - PubMed Abstract:
Bacteriorhodopsin (BR) and sensory rhodopsin II (SRII), homologous photoactive proteins in haloarchaea, have different molecular functions. BR is a light-driven proton pump, whereas SRII is a phototaxis receptor that transmits a light-induced conformational change to its transducer HtrII. Despite these distinctly different functions, a single residue substitution, Ala215 to Thr215 in the BR retinal-binding pocket, enables its photochemical reactions to transmit signals to HtrII and mediate phototaxis. We pursued a crystal structure of the signaling BR mutant (BR_A215T) to determine the structural changes caused by the A215T mutation and to assess what new photochemistry is likely to be introduced into the BR photoactive site. We crystallized BR_A215T from bicelles and solved its structure to 3.0 Å resolution to enable an atomic-level comparison. The analysis was complemented by molecular dynamics simulation of BR mutated in silico. Three main conclusions regarding the roles of photoactive site residues in signaling emerge from the comparison of BR_A215T, BR, and SRII structures: (i) the Thr215 residue in signaling BR is positioned nearly identically with respect to the retinal chromophore as in SRII, consistent with its role in producing a steric conflict with the retinal C₁₄ group during photoisomerization, proposed earlier to be essential for SRII signaling from vibrational spectroscopy and motility measurements; (ii) Tyr174-Thr204 hydrogen bonding, critical in SRII signaling and mimicked in signaling BR, is likely auxiliary, for example, to maintain Thr204 in the proper position for the steric trigger to occur; and (iii) the primary role of Arg72 in SRII is spectral tuning and not signaling.
- Department of Biochemistry and Molecular Biology, Center for Membrane Biology, University of Texas Medical School, Houston, TX 77030, USA. john.l.spudich@uth.tmc.edu
Organizational Affiliation: