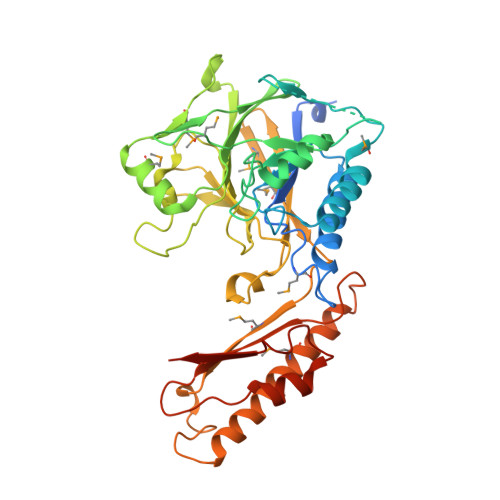
Crystal structure of human mre11: understanding tumorigenic mutations
Park, Y.B., Chae, J., Kim, Y., Cho, Y.(2011) Structure 19: 1591-1602
- PubMed: 22078559 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2011.09.010
- Primary Citation Related Structures:
3T1I - PubMed Abstract:
Mre11 plays an important role in repairing damaged DNA by cleaving broken ends and by providing a platform for other DNA repair proteins. Various Mre11 mutations have been identified in several types of cancer. We have determined the crystal structure of the human Mre11 core (hMre11), which contains the nuclease and capping domains. hMre11 dimerizes through the interfaces between loop β3-α3 from one Mre11 and loop β4-β5 from another Mre11, and between loop α2-β3 from one Mre11 and helices α2 and α3 from another Mre11, and assembles into a completely different dimeric architecture compared with bacterial or archaeal Mre11 homologs. Nbs1 binds to the region containing loop α2-β3 which participates in dimerization. The hMre11 structure in conjunction with biochemical analyses reveals that many tumorigenic mutations are primarily associated with Nbs1 binding and partly with nuclease activities, providing a framework for understanding how mutations inactivate Mre11.
- Department of Life Science, Pohang University of Science and Technology, Pohang 790-784, South Korea.
Organizational Affiliation: