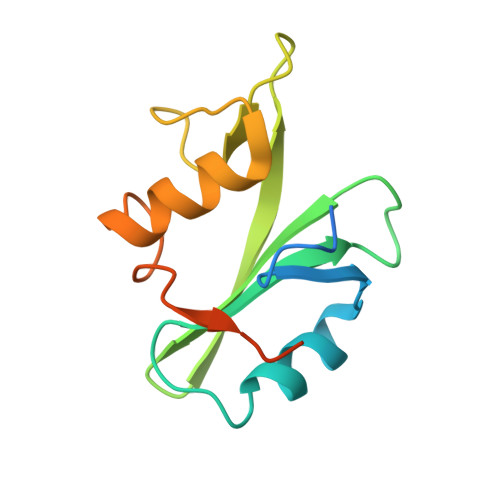
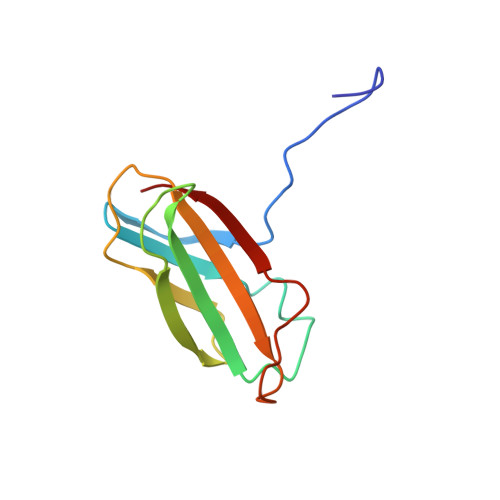
Targeting the SH2-Kinase Interface in Bcr-Abl Inhibits Leukemogenesis.
Grebien, F., Hantschel, O., Wojcik, J., Kaupe, I., Kovacic, B., Wyrzucki, A.M., Gish, G.D., Cerny-Reiterer, S., Koide, A., Beug, H., Pawson, T., Valent, P., Koide, S., Superti-Furga, G.(2011) Cell 147: 306-319
- PubMed: 22000011 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2011.08.046
- Primary Citation Related Structures:
3T04 - PubMed Abstract:
Chronic myelogenous leukemia (CML) is caused by the constitutively active tyrosine kinase Bcr-Abl and treated with the tyrosine kinase inhibitor (TKI) imatinib. However, emerging TKI resistance prevents complete cure. Therefore, alternative strategies targeting regulatory modules of Bcr-Abl in addition to the kinase active site are strongly desirable. Here, we show that an intramolecular interaction between the SH2 and kinase domains in Bcr-Abl is both necessary and sufficient for high catalytic activity of the enzyme. Disruption of this interface led to inhibition of downstream events critical for CML signaling and, importantly, completely abolished leukemia formation in mice. Furthermore, disruption of the SH2-kinase interface increased sensitivity of imatinib-resistant Bcr-Abl mutants to TKI inhibition. An engineered Abl SH2-binding fibronectin type III monobody inhibited Bcr-Abl kinase activity both in vitro and in primary CML cells, where it induced apoptosis. This work validates the SH2-kinase interface as an allosteric target for therapeutic intervention.
- Research Center for Molecular Medicine of the Austrian Academy of Sciences, Vienna, Austria.
Organizational Affiliation: