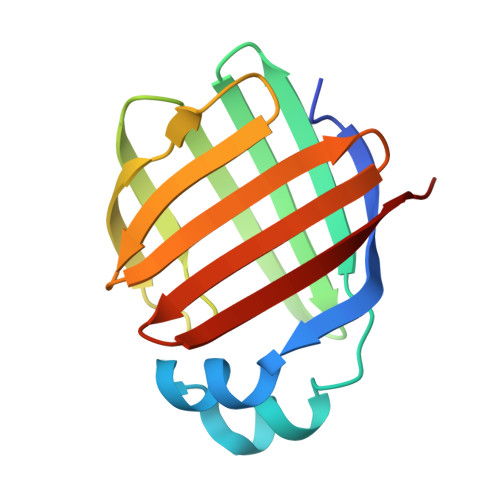
Fatty acid induced remodeling within the Human liver fatty acid binding protein
Sharma, A., Sharma, A.(2011) J Biological Chem
- PubMed: 21757748
- DOI: https://doi.org/10.1074/jbc.M111.270165
- Primary Citation Related Structures:
3STK, 3STM, 3STN - PubMed Abstract:
We crystallized human liver fatty acid-binding protein (LFABP) in apo, holo, and intermediate states of palmitic acid engagement. Structural snapshots of fatty acid recognition, entry, and docking within LFABP support a heads-in mechanism for ligand entry. Apo-LFABP undergoes structural remodeling, where the first palmitate ingress creates the atomic environment for placement of the second palmitate. These new mechanistic insights will facilitate development of pharmacological agents against LFABP.
- Structural and Computational Biology Group, International Centre for Genetic Engineering and Biotechnology, Aruna Asaf Ali Road, 110067 New Delhi, India.
Organizational Affiliation: