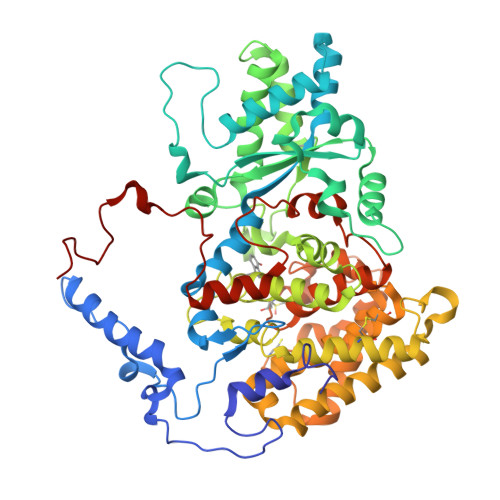
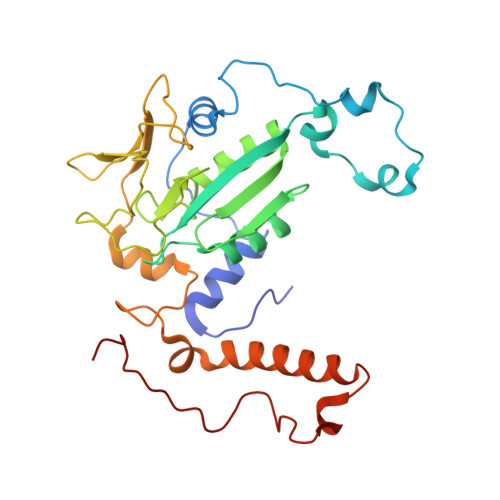
Structure of a methyl-coenzyme M reductase from Black Sea mats that oxidize methane anaerobically.
Shima, S., Krueger, M., Weinert, T., Demmer, U., Kahnt, J., Thauer, R.K., Ermler, U.(2011) Nature 481: 98-101
- PubMed: 22121022 Search on PubMed
- DOI: https://doi.org/10.1038/nature10663
- Primary Citation Related Structures:
3SQG - PubMed Abstract:
The anaerobic oxidation of methane (AOM) with sulphate, an area currently generating great interest in microbiology, is accomplished by consortia of methanotrophic archaea (ANME) and sulphate-reducing bacteria. The enzyme activating methane in methanotrophic archaea has tentatively been identified as a homologue of methyl-coenzyme M reductase (MCR) that catalyses the methane-forming step in methanogenic archaea. Here we report an X-ray structure of the 280 kDa heterohexameric ANME-1 MCR complex. It was crystallized uniquely from a protein ensemble purified from consortia of microorganisms collected with a submersible from a Black Sea mat catalysing AOM with sulphate. Crystals grown from the heterogeneous sample diffract to 2.1 Å resolution and consist of a single ANME-1 MCR population, demonstrating the strong selective power of crystallization. The structure revealed ANME-1 MCR in complex with coenzyme M and coenzyme B, indicating the same substrates for MCR from methanotrophic and methanogenic archaea. Differences between the highly similar structures of ANME-1 MCR and methanogenic MCR include a F(430) modification, a cysteine-rich patch and an altered post-translational amino acid modification pattern, which may tune the enzymes for their functions in different biological contexts.
- Max Planck Institute for Terrestrial Microbiology, Karl-Frisch-Strasse 10, D-35043 Marburg, Germany. shima@mpi-marburg.mpg.de
Organizational Affiliation: