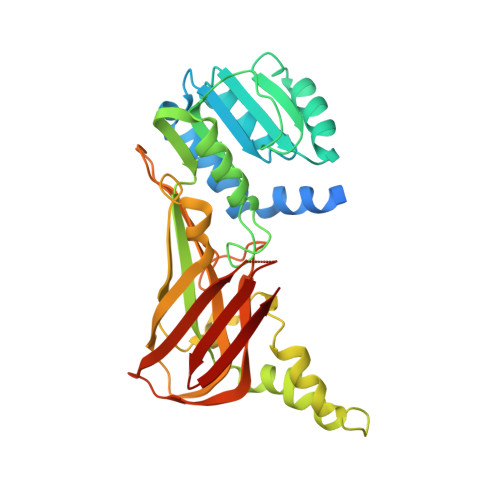
An allosteric inhibitor of protein arginine methyltransferase 3.
Siarheyeva, A., Senisterra, G., Allali-Hassani, A., Dong, A., Dobrovetsky, E., Wasney, G.A., Chau, I., Marcellus, R., Hajian, T., Liu, F., Korboukh, I., Smil, D., Bolshan, Y., Min, J., Wu, H., Zeng, H., Loppnau, P., Poda, G., Griffin, C., Aman, A., Brown, P.J., Jin, J., Al-Awar, R., Arrowsmith, C.H., Schapira, M., Vedadi, M.(2012) Structure 20: 1425-1435
- PubMed: 22795084 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.06.001
- Primary Citation Related Structures:
3SMQ - PubMed Abstract:
PRMT3, a protein arginine methyltransferase, has been shown to influence ribosomal biosynthesis by catalyzing the dimethylation of the 40S ribosomal protein S2. Although PRMT3 has been reported to be a cytosolic protein, it has been shown to methylate histone H4 peptide (H4 1-24) in vitro. Here, we report the identification of a PRMT3 inhibitor (1-(benzo[d][1,2,3]thiadiazol-6-yl)-3-(2-cyclohexenylethyl)urea; compound 1) with IC50 value of 2.5 μM by screening a library of 16,000 compounds using H4 (1-24) peptide as a substrate. The crystal structure of PRMT3 in complex with compound 1 as well as kinetic analysis reveals an allosteric mechanism of inhibition. Mutating PRMT3 residues within the allosteric site or using compound 1 analogs that disrupt interactions with allosteric site residues both abrogated binding and inhibitory activity. These data demonstrate an allosteric mechanism for inhibition of protein arginine methyltransferases, an emerging class of therapeutic targets.
- Structural Genomics Consortium, University of Toronto, 101 College Street, MaRS Centre, South Tower, Toronto, ON M5G 1L7, Canada.
Organizational Affiliation: