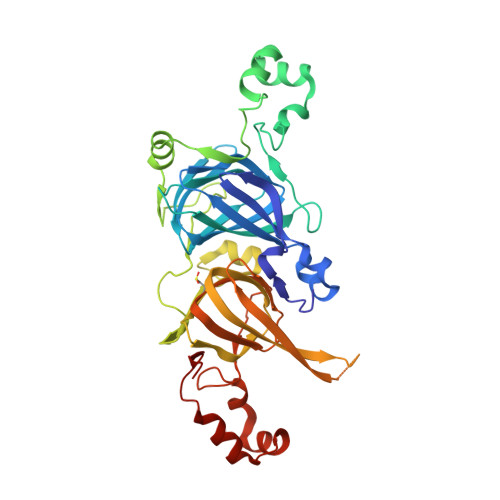
Crystal structure of the major peanut allergen Ara h 1.
Cabanos, C., Urabe, H., Tandang-Silvas, M.R., Utsumi, S., Mikami, B., Maruyama, N.(2011) Mol Immunol 49: 115-123
- PubMed: 21903274 Search on PubMed
- DOI: https://doi.org/10.1016/j.molimm.2011.08.004
- Primary Citation Related Structures:
3SMH - PubMed Abstract:
Ara h 1, a 7S globulin, is one of the three major peanut allergens. We previously reported the crystallization of the core region of recombinant Ara h 1. Here, we present the crystal structure of the Ara h 1 core at a resolution of 2.43 Å. We also assayed the Ara h 1 core thermal stability and compared its final structure against other 7S globulins. The Ara h 1 core has a thermal denaturation temperature of 88.3°C and a structure that is very similar to other 7S globulins. Previously identified linear IgE epitopes were also mapped on the three-dimensional structure. Most linear epitopes were found in the extended loop domains and the coils between the N- and C-terminal modules, while others were found in the less accessible β-sheets of the C-terminal core β-barrel domain of each monomer. Most of these epitopes become either slightly or significantly buried upon trimer formation, implying that allergen digestion in the gut is required for these epitopes to be accessible to immunoglobulins. Our findings also suggest that both intact and partially degraded allergens should be employed in future diagnostic and immunotherapeutic strategies.
- Laboratory of Food Quality Design and Development, Graduate School of Agriculture, Kyoto University Uji, Kyoto, Japan.
Organizational Affiliation: