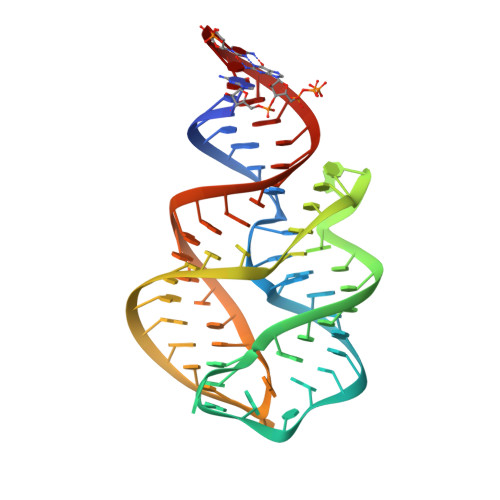
Structural principles of nucleoside selectivity in a 2'-deoxyguanosine riboswitch.
Pikovskaya, O., Polonskaia, A., Patel, D.J., Serganov, A.(2011) Nat Chem Biol 7: 748-755
- PubMed: 21841796 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.631
- Primary Citation Related Structures:
3SKI, 3SKL, 3SKR, 3SKT, 3SKW, 3SKZ, 3SLM, 3SLQ - PubMed Abstract:
Purine riboswitches have an essential role in genetic regulation of bacterial metabolism. This family includes the 2'-deoxyguanosine (dG) riboswitch, which is involved in feedback control of deoxyguanosine biosynthesis. To understand the principles that define dG selectivity, we determined crystal structures of the natural Mesoplasma florum riboswitch bound to cognate dG as well as to noncognate guanosine, deoxyguanosine monophosphate and guanosine monophosphate. Comparison with related purine riboswitch structures reveals that the dG riboswitch achieves its specificity through modification of key interactions involving the nucleobase and rearrangement of the ligand-binding pocket to accommodate the additional sugar moiety. In addition, we observe new conformational changes beyond the junctional binding pocket extending as far as peripheral loop-loop interactions. It appears that re-engineering riboswitch scaffolds will require consideration of selectivity features dispersed throughout the riboswitch tertiary fold, and structure-guided drug design efforts targeted to junctional RNA scaffolds need to be addressed within such an expanded framework.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York, USA.
Organizational Affiliation: