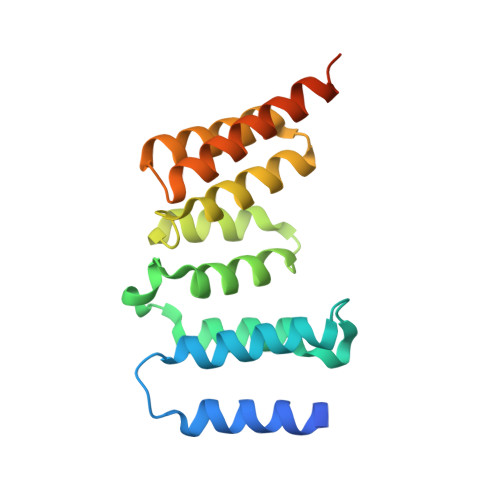
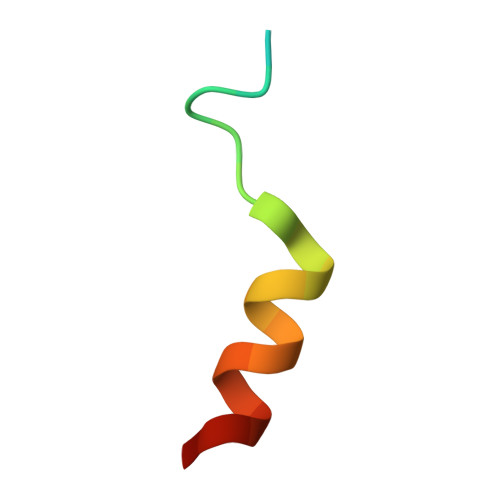
Structure of a Blinkin-BUBR1 Complex Reveals an Interaction Crucial for Kinetochore-Mitotic Checkpoint Regulation via an Unanticipated Binding Site.
Bolanos-Garcia, V.M., Lischetti, T., Matak-Vinkovic, D., Cota, E., Simpson, P.J., Chirgadze, D.Y., Spring, D.R., Robinson, C.V., Nilsson, J., Blundell, T.L.(2011) Structure 19: 1691-1700
- PubMed: 22000412 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.09.017
- Primary Citation Related Structures:
3SI5 - PubMed Abstract:
The maintenance of genomic stability relies on the spindle assembly checkpoint (SAC), which ensures accurate chromosome segregation by delaying the onset of anaphase until all chromosomes are properly bioriented and attached to the mitotic spindle. BUB1 and BUBR1 kinases are central for this process and by interacting with Blinkin, link the SAC with the kinetochore, the macromolecular assembly that connects microtubules with centromeric DNA. Here, we identify the Blinkin motif critical for interaction with BUBR1, define the stoichiometry and affinity of the interaction, and present a 2.2 Å resolution crystal structure of the complex. The structure defines an unanticipated BUBR1 region responsible for the interaction and reveals a novel Blinkin motif that undergoes a disorder-to-order transition upon ligand binding. We also show that substitution of several BUBR1 residues engaged in binding Blinkin leads to defects in the SAC, thus providing the first molecular details of the recognition mechanism underlying kinetochore-SAC signaling.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, UK. victor@cryst.bioc.cam.ac.uk
Organizational Affiliation: