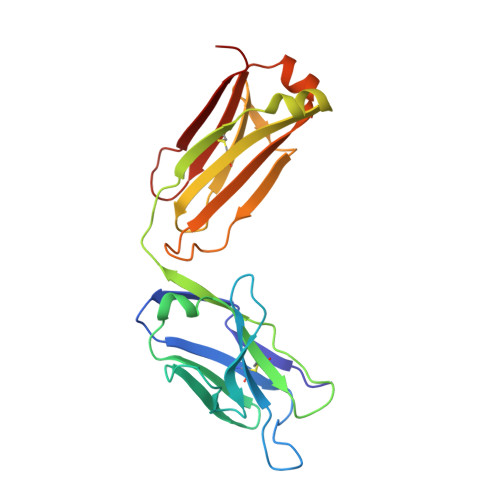
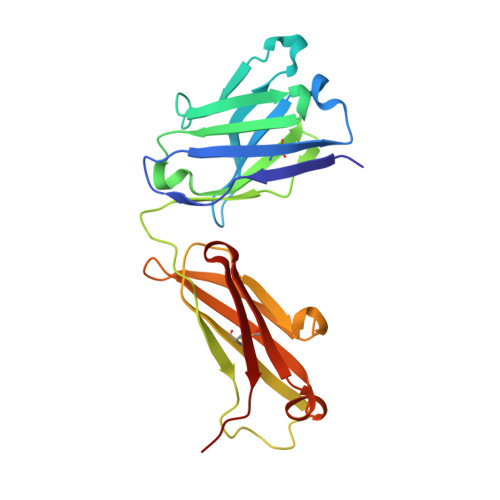
Crystal structure of the complex mAb 17.2 and the C-terminal region of Trypanosoma cruzi P2 Beta protein: implications in cross-reactivity
Pizarro, J.C., Boulot, G., Bentley, G.A., Gomez, K.A., Hoebeke, J., Hontebeyrie, M., Levin, M.J., Smulski, C.R.(2011) PLoS Negl Trop Dis 5: e1375-e1375
- PubMed: 22069505 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pntd.0001375
- Primary Citation Related Structures:
3SGD, 3SGE - PubMed Abstract:
Patients with Chronic Chagas' Heart Disease possess high levels of antibodies against the carboxyl-terminal end of the ribosomal P2ß protein of Trypanosoma cruzi (TcP2ß). These antibodies, as well as the murine monoclonal antibody (mAb) 17.2, recognize the last 13 amino acids of TcP2ß (called the R13 epitope: EEEDDDMGFGLFD) and are able to cross-react with, and stimulate, the ß1 adrenergic receptor (ß1-AR). Indeed, the mAb 17.2 was able to specifically detect human β1-AR, stably transfected into HEK cells, by flow cytometry and to induce repolarisation abnormalities and first degree atrioventricular conduction block after passive transfer to naïve mice. To study the structural basis of this cross-reactivity, we determined the crystal structure of the Fab region of the mAb 17.2 alone at 2.31 Å resolution and in complex with the R13 peptide at 1.89 Å resolution. We identified as key contact residues on R13 peptide Glu3, Asp6 and Phe9 as was previously shown by alanine scanning. Additionally, we generated a model of human β1-AR to elucidate the interaction with anti-R13 antibodies. These data provide an understanding of the molecular basis of cross-reactive antibodies induced by chronic infection with Trypanosoma cruzi.
- Unité d'Immunologie Structurale, Institut Pasteur, Paris, France.
Organizational Affiliation: