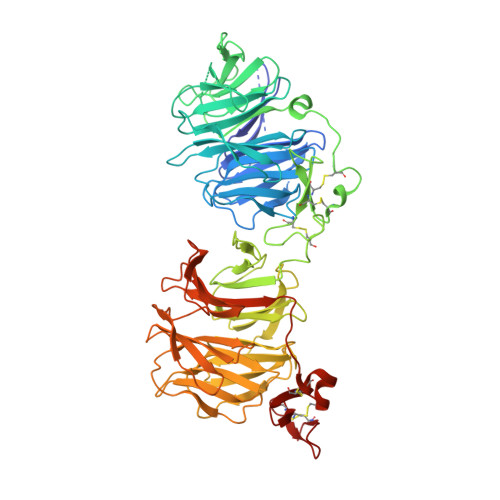
Crystal structures of the extracellular domain of LRP6 and its complex with DKK1.
Cheng, Z., Biechele, T., Wei, Z., Morrone, S., Moon, R.T., Wang, L., Xu, W.(2011) Nat Struct Mol Biol 18: 1204-1210
- PubMed: 21984209 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2139
- Primary Citation Related Structures:
3S8V, 3S8Z, 3S94 - PubMed Abstract:
Low-density-lipoprotein (LDL) receptor-related proteins 5 and 6 (LRP5/6) are Wnt co-receptors essential for Wnt/β-catenin signaling. Dickkopf 1 (DKK1) inhibits Wnt signaling by interacting with the extracellular domains of LRP5/6 and is a drug target for multiple diseases. Here we present the crystal structures of a human LRP6-E3E4-DKK1 complex and the first and second halves of human LRP6's four propeller-epidermal growth factor (EGF) pairs (LRP6-E1E2 and LRP6-E3E4). Combined with EM analysis, these data demonstrate that LRP6-E1E2 and LRP6-E3E4 form two rigid structural blocks, with a short intervening hinge that restrains their relative orientation. The C-terminal domain of DKK1 (DKK1c) interacts with the top surface of the LRP6-E3 YWTD propeller and given their structural similarity, probably also that of the LRP6-E1 propeller, through conserved hydrophobic patches buttressed by a network of salt bridges and hydrogen bonds. Our work provides key insights for understanding LRP5/6 structure and the interaction of LRP5/6 with DKK, as well as for drug discovery.
- Department of Biological Structure, University of Washington School of Medicine, Seattle, Washington, USA.
Organizational Affiliation: