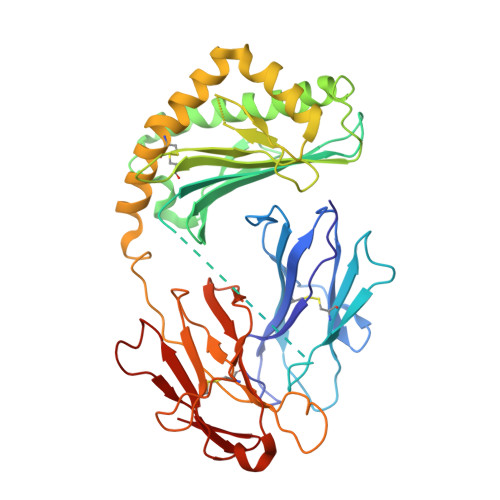
Crystal structure of human CD1e reveals a groove suited for lipid-exchange processes.
Garcia-Alles, L.F., Giacometti, G., Versluis, C., Maveyraud, L., de Paepe, D., Guiard, J., Tranier, S., Gilleron, M., Prandi, J., Hanau, D., Heck, A.J., Mori, L., De Libero, G., Puzo, G., Mourey, L., de la Salle, H.(2011) Proc Natl Acad Sci U S A 108: 13230-13235
- PubMed: 21788486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1105627108
- Primary Citation Related Structures:
3S6C - PubMed Abstract:
CD1e is the only human CD1 protein existing in soluble form in the late endosomes of dendritic cells, where it facilitates the processing of glycolipid antigens that are ultimately recognized by CD1b-restricted T cells. The precise function of CD1e remains undefined, thus impeding efforts to predict the participation of this protein in the presentation of other antigens. To gain insight into its function, we determined the crystal structure of recombinant CD1e expressed in human cells at 2.90-Å resolution. The structure revealed a groove less intricate than in other CD1 proteins, with a significantly wider portal characterized by a 2 Å-larger spacing between the α1 and α2 helices. No electron density corresponding to endogenous ligands was detected within the groove, despite the presence of ligands unequivocally established by native mass spectrometry in recombinant CD1e. Our structural data indicate that the water-exposed CD1e groove could ensure the establishment of loose contacts with lipids. In agreement with this possibility, lipid association and dissociation processes were found to be considerably faster with CD1e than with CD1b. Moreover, CD1e was found to mediate in vitro the transfer of lipids to CD1b and the displacement of lipids from stable CD1b-antigen complexes. Altogether, these data support that CD1e could have evolved to mediate lipid-exchange/editing processes with CD1b and point to a pathway whereby the repertoire of lipid antigens presented by human dendritic cells might be expanded.
- Centre National de la Recherche Scientifique, Institut de Pharmacologie et de Biologie Structurale, Toulouse F-31077, France.
Organizational Affiliation: