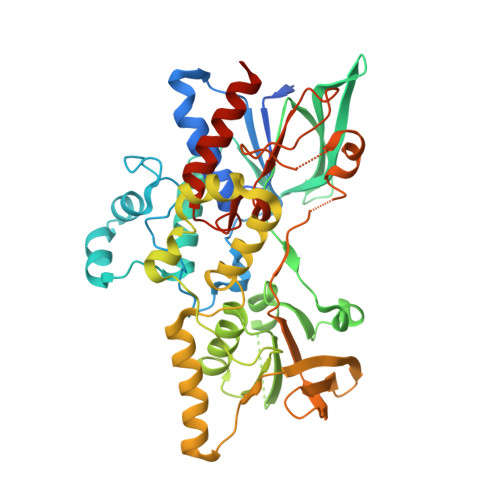
Two Structures of an N-Hydroxylating Flavoprotein Monooxygenase: ORNITHINE HYDROXYLASE FROM PSEUDOMONAS AERUGINOSA.
Olucha, J., Meneely, K.M., Chilton, A.S., Lamb, A.L.(2011) J Biological Chem 286: 31789-31798
- PubMed: 21757711
- DOI: https://doi.org/10.1074/jbc.M111.265876
- Primary Citation of Related Structures:
3S5W, 3S61 - PubMed Abstract:
The ornithine hydroxylase from Pseudomonas aeruginosa (PvdA) catalyzes the FAD-dependent hydroxylation of the side chain amine of ornithine, which is subsequently formylated to generate the iron-chelating hydroxamates of the siderophore pyoverdin. PvdA belongs to the class B flavoprotein monooxygenases, which catalyze the oxidation of substrates using NADPH as the electron donor and molecular oxygen. Class B enzymes include the well studied flavin-containing monooxygenases and Baeyer-Villiger monooxygenases. The first two structures of a class B N-hydroxylating monooxygenase were determined with FAD in oxidized (1.9 Å resolution) and reduced (3.03 Å resolution) states. PvdA has the two expected Rossmann-like dinucleotide-binding domains for FAD and NADPH and also a substrate-binding domain, with the active site at the interface between the three domains. The structures have NADP(H) and (hydroxy)ornithine bound in a solvent-exposed active site, providing structural evidence for substrate and co-substrate specificity and the inability of PvdA to bind FAD tightly. Structural and biochemical evidence indicates that NADP(+) remains bound throughout the oxidative half-reaction, which is proposed to shelter the flavin intermediates from solvent and thereby prevent uncoupling of NADPH oxidation from hydroxylated product formation.
- Department of Molecular Biosciences, University of Kansas, Lawrence, Kansas 66045, USA.
Organizational Affiliation: