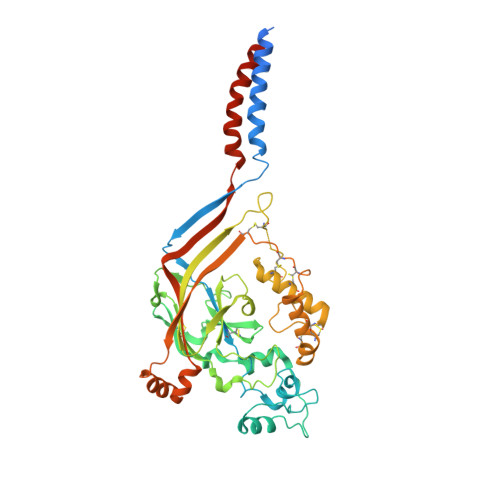
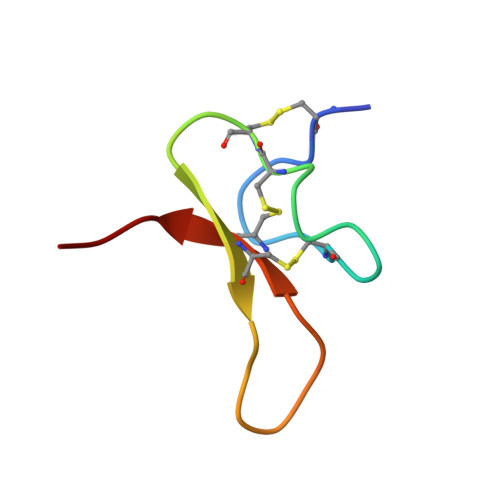
Structure of the Acid-sensing ion channel 1 in complex with the gating modifier Psalmotoxin 1.
Dawson, R.J., Benz, J., Stohler, P., Tetaz, T., Joseph, C., Huber, S., Schmid, G., Hugin, D., Pflimlin, P., Trube, G., Rudolph, M.G., Hennig, M., Ruf, A.(2012) Nat Commun 3: 936-936
- PubMed: 22760635 Search on PubMed
- DOI: https://doi.org/10.1038/ncomms1917
- Primary Citation Related Structures:
3S3W, 3S3X - PubMed Abstract:
Venom-derived peptide toxins can modify the gating characteristics of excitatory channels in neurons. How they bind and interfere with the flow of ions without directly blocking the ion permeation pathway remains elusive. Here we report the crystal structure of the trimeric chicken Acid-sensing ion channel 1 in complex with the highly selective gating modifier Psalmotoxin 1 at 3.0 Å resolution. The structure reveals the molecular interactions of three toxin molecules binding at the proton-sensitive acidic pockets of Acid-sensing ion channel 1 and electron density consistent with a cation trapped in the central vestibule above the ion pathway. A hydrophobic patch and a basic cluster are the key structural elements of Psalmotoxin 1 binding, locking two separate regulatory regions in their relative, desensitized-like arrangement. Our results provide a general concept for gating modifier toxin binding suggesting that both surface motifs are required to modify the gating characteristics of an ion channel.
- F. Hoffmann-La Roche AG, pRED, Pharma Research & Early Development, Discovery Technologies, Grenzacherstrasse 124, Basel CH4070, Switzerland. roger.dawson@roche.com
Organizational Affiliation: