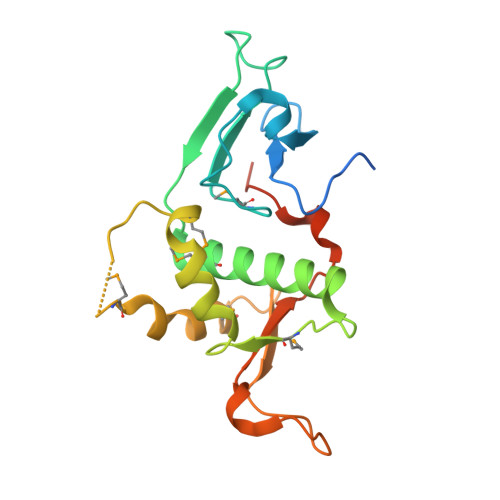
Crystal structure of the trithorax group protein ASH2L reveals a forkhead-like DNA binding domain.
Sarvan, S., Avdic, V., Tremblay, V., Chaturvedi, C.P., Zhang, P., Lanouette, S., Blais, A., Brunzelle, J.S., Brand, M., Couture, J.F.(2011) Nat Struct Mol Biol 18: 857-859
- PubMed: 21642971 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2093
- Primary Citation Related Structures:
3S32 - PubMed Abstract:
Absent, small or homeotic discs-like 2 (ASH2L) is a trithorax group (TrxG) protein and a regulatory subunit of the SET1 family of lysine methyltransferases. Here we report that ASH2L binds DNA using a forkhead-like helix-wing-helix (HWH) domain. In vivo, the ASH2L HWH domain is required for binding to the β-globin locus control region, histone H3 Lys4 (H3K4) trimethylation and maximal expression of the β-globin gene (Hbb-1), validating the functional importance of the ASH2L DNA binding domain.
- Ottawa Institute of Systems Biology, Department of Biochemistry, University of Ottawa, Ottawa, Ontario, Canada.
Organizational Affiliation: