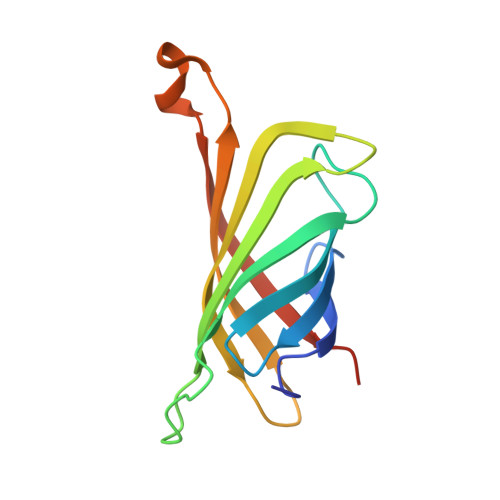
Streptavidin and its biotin complex at atomic resolution.
Le Trong, I., Wang, Z., Hyre, D.E., Lybrand, T.P., Stayton, P.S., Stenkamp, R.E.(2011) Acta Crystallogr D Biol Crystallogr 67: 813-821
- PubMed: 21904034 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911027806
- Primary Citation Related Structures:
3RY1, 3RY2 - PubMed Abstract:
Atomic resolution crystallographic studies of streptavidin and its biotin complex have been carried out at 1.03 and 0.95 Å, respectively. The wild-type protein crystallized with a tetramer in the asymmetric unit, while the crystals of the biotin complex contained two subunits in the asymmetric unit. Comparison of the six subunits shows the various ways in which the protein accommodates ligand binding and different crystal-packing environments. Conformational variation is found in each of the polypeptide loops connecting the eight strands in the β-sandwich subunit, but the largest differences are found in the flexible binding loop (residues 45-52). In three of the unliganded subunits the loop is in an `open' conformation, while in the two subunits binding biotin, as well as in one of the unliganded subunits, this loop `closes' over the biotin-binding site. The `closed' loop contributes to the protein's high affinity for biotin. Analysis of the anisotropic displacement parameters included in the crystallographic models is consistent with the variation found in the loop structures and the view that the dynamic nature of the protein structure contributes to the ability of the protein to bind biotin so tightly.
- Department of Biological Structure, University of Washington, Seattle, USA.
Organizational Affiliation: