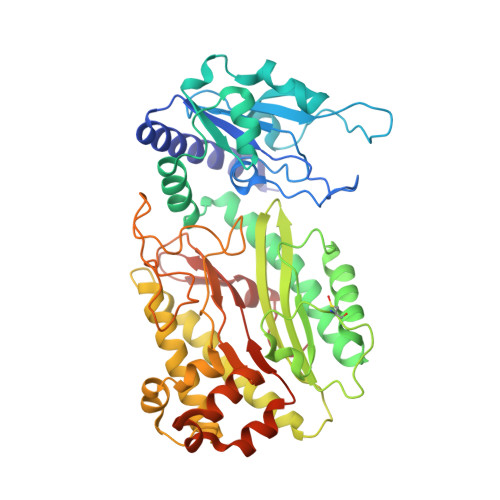
Organophosphorus acid anhydrolase from Alteromonas macleodii: structural study and functional relationship to prolidases.
Stepankova, A., Duskova, J., Skalova, T., Hasek, J., Koval, T., Ostergaard, L.H., Dohnalek, J.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 346-354
- PubMed: 23545636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113002674
- Primary Citation Related Structures:
3RVA - PubMed Abstract:
The bacterial enzyme organophosphorus acid anhydrolase (OPAA) is able to catalyze the hydrolysis of both proline dipeptides (Xaa-Pro) and several types of organophosphate (OP) compounds. The full three-dimensional structure of the manganese-dependent OPAA enzyme is presented for the first time. This enzyme, which was originally isolated from the marine bacterium Alteromonas macleodii, was prepared recombinantly in Escherichia coli. The crystal structure was determined at 1.8 Å resolution in space group C2, with unit-cell parameters a = 133.8, b = 49.2, c = 97.3 Å, β = 125.0°. The enzyme forms dimers and their existence in solution was confirmed by dynamic light scattering and size-exclusion chromatography. The enzyme shares the pita-bread fold of its C-terminal domain with related prolidases. The binuclear manganese centre is located in the active site within the pita-bread domain. Moreover, an Ni(2+) ion from purification was localized according to anomalous signal. This study presents the full structure of this enzyme with complete surroundings of the active site and provides a critical analysis of its relationship to prolidases.
- Institute of Macromolecular Chemistry, AS CR, vvi, Heyrovsky sq 2, 162 06 Prague 6, Czech Republic. a.stepanko@gmail.com
Organizational Affiliation: