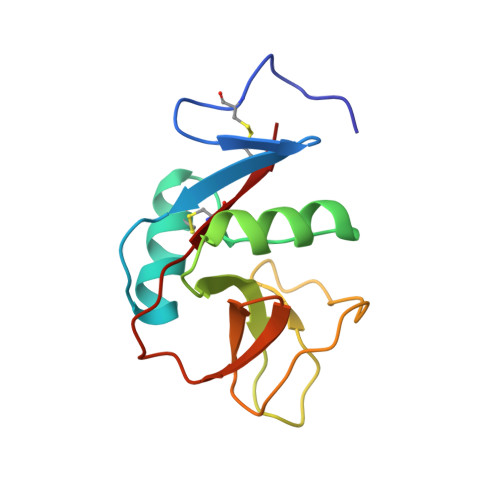
Mouse Clr-g, a Ligand for NK Cell Activation Receptor NKR-P1F: Crystal Structure and Biophysical Properties.
Skalova, T., Kotynkova, K., Duskova, J., Hasek, J., Kovai, T., Kolenko, P., Novak, P., Man, P., Hanc, P., Vanek, O., Bezouska, K., Dohnalek, J.(2012) J Immunol 189: 4881-4889
- PubMed: 23071282 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.1200880
- Primary Citation Related Structures:
3RS1 - PubMed Abstract:
Interactions between C-type lectin-like NK cell receptors and their protein ligands form one of the key recognition mechanisms of the innate immune system that is involved in the elimination of cells that have been malignantly transformed, virally infected, or stressed by chemotherapy or other factors. We determined an x-ray structure for the extracellular domain of mouse C-type lectin related (Clr) protein g, a ligand for the activation receptor NKR-P1F. Clr-g forms dimers in the crystal structure resembling those of human CD69. This newly reported structure, together with the previously determined structure of mouse receptor NKR-P1A, allowed the modeling and calculations of electrostatic profiles for other closely related receptors and ligands. Despite the high similarity among Clr-g, Clr-b, and human CD69, these molecules have fundamentally different electrostatics, with distinct polarization of Clr-g. The electrostatic profile of NKR-P1F is complementary to that of Clr-g, which suggests a plausible interaction mechanism based on contacts between surface sites of opposite potential.
- Institute of Macromolecular Chemistry, Academy of Sciences of the Czech Republic, vvi, 16206 Praha 6, Czech Republic. t.skalova@gmail.com
Organizational Affiliation: