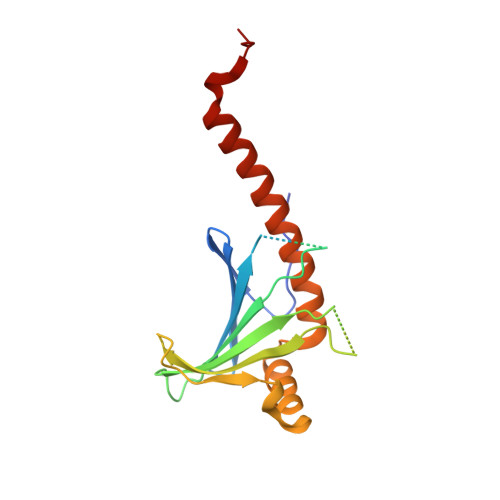
Crystal structure of complement component 1, q subcomponent binding protein, C1QBP
Tempel, W., Tong, Y., Crombet, L., Shen, Y., Guan, X., Nedyalkova, L., Walker, J.R., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Weigelt, J., Park, H.To be published.