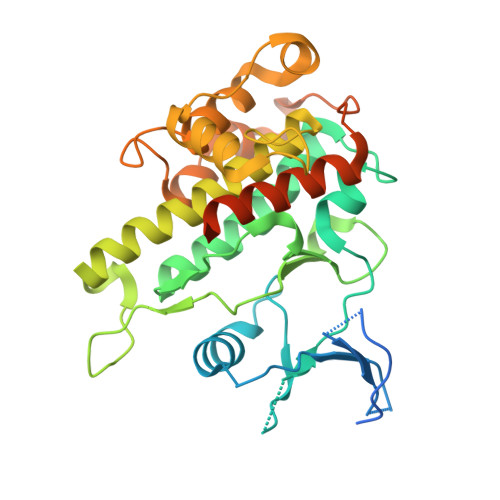
Structural basis for the autoinhibition of the C-terminal kinase domain of human RSK1.
Li, D., Fu, T.M., Nan, J., Liu, C., Li, L.F., Su, X.D.(2012) Acta Crystallogr D Biol Crystallogr 68: 680-685
- PubMed: 22683790 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912007457
- Primary Citation Related Structures:
3RNY - PubMed Abstract:
p90 ribosomal S6 kinases (RSKs) respond to various mitogen stimuli and comprise two distinct protein kinase domains. The C-terminal kinase domain (CTKD) receives signal from ERK1/2 and adopts an autoinhibitory mechanism. Here, the crystal structure of human RSK1 CTKD is reported at 2.7 Å resolution. The structure shows a standard kinase fold, with the catalytic residues in the ATP-binding cleft orientated in optimal conformations for phosphotransfer. The inactivation of the CTKD is conferred by an extra α-helix (αL), which occupies the substrate-binding groove. In combination with previous knowledge, this structure indicates that activation of RSK1 involves the removal of αL from the substrate-binding groove induced by ERK1/2 phosphorylation.
- State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing 100871, People's Republic of China.
Organizational Affiliation: