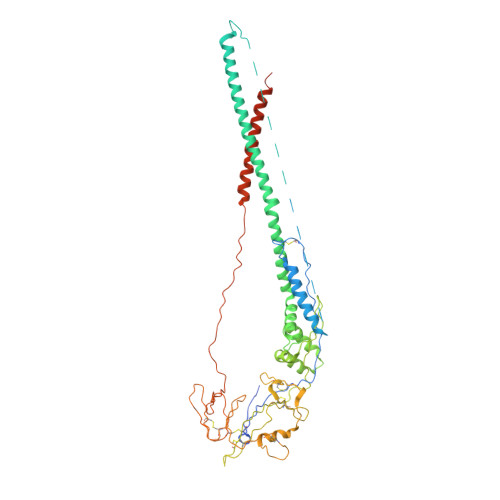
Structural basis for immunization with postfusion respiratory syncytial virus fusion F glycoprotein (RSV F) to elicit high neutralizing antibody titers.
Swanson, K.A., Settembre, E.C., Shaw, C.A., Dey, A.K., Rappuoli, R., Mandl, C.W., Dormitzer, P.R., Carfi, A.(2011) Proc Natl Acad Sci U S A 108: 9619-9624
- PubMed: 21586636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1106536108
- Primary Citation Related Structures:
3RKI - PubMed Abstract:
Respiratory syncytial virus (RSV), the main cause of infant bronchiolitis, remains a major unmet vaccine need despite more than 40 years of vaccine research. Vaccine candidates based on a chief RSV neutralization antigen, the fusion (F) glycoprotein, have foundered due to problems with stability, purity, reproducibility, and potency. Crystal structures of related parainfluenza F glycoproteins have revealed a large conformational change between the prefusion and postfusion states, suggesting that postfusion F antigens might not efficiently elicit neutralizing antibodies. We have generated a homogeneous, stable, and reproducible postfusion RSV F immunogen that elicits high titers of neutralizing antibodies in immunized animals. The 3.2-Å X-ray crystal structure of this substantially complete RSV F reveals important differences from homology-based structural models. Specifically, the RSV F crystal structure demonstrates the exposure of key neutralizing antibody binding sites on the surface of the postfusion RSV F trimer. This unanticipated structural feature explains the engineered RSV F antigen's efficiency as an immunogen. This work illustrates how structural-based antigen design can guide the rational optimization of candidate vaccine antigens.
- Novartis Vaccines and Diagnostics, Cambridge, MA 02139, USA.
Organizational Affiliation: