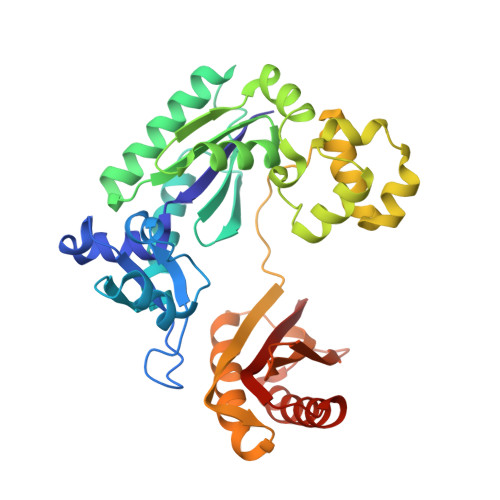
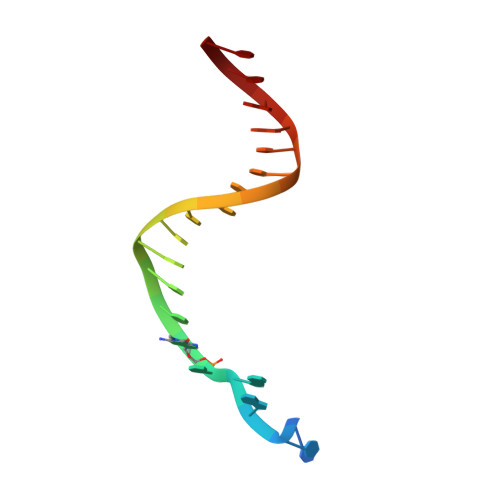
Implications for damage recognition during Dpo4-mediated mutagenic bypass of m1G and m3C lesions.
Rechkoblit, O., Delaney, J.C., Essigmann, J.M., Patel, D.J.(2011) Structure 19: 821-832
- PubMed: 21645853 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.03.020
- Primary Citation Related Structures:
3RAQ, 3RAX, 3RB0, 3RB3, 3RB4, 3RB6, 3RBD, 3RBE - PubMed Abstract:
DNA is susceptible to alkylation damage by a number of environmental agents that modify the Watson-Crick edge of the bases. Such lesions, if not repaired, may be bypassed by Y-family DNA polymerases. The bypass polymerase Dpo4 is strongly inhibited by 1-methylguanine (m1G) and 3-methylcytosine (m3C), with nucleotide incorporation opposite these lesions being predominantly mutagenic. Further, extension after insertion of both correct and incorrect bases, introduces additional base substitution and deletion errors. Crystal structures of the Dpo4 ternary extension complexes with correct and mismatched 3'-terminal primer bases opposite the lesions reveal that both m1G and m3C remain positioned within the DNA template/primer helix. However, both correct and incorrect pairing partners exhibit pronounced primer terminal nucleotide distortion, being primarily evicted from the DNA helix when opposite m1G or misaligned when pairing with m3C. Our studies provide insights into mechanisms related to hindered and mutagenic bypass of methylated lesions and models associated with damage recognition by repair demethylases.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10021, USA.
Organizational Affiliation: