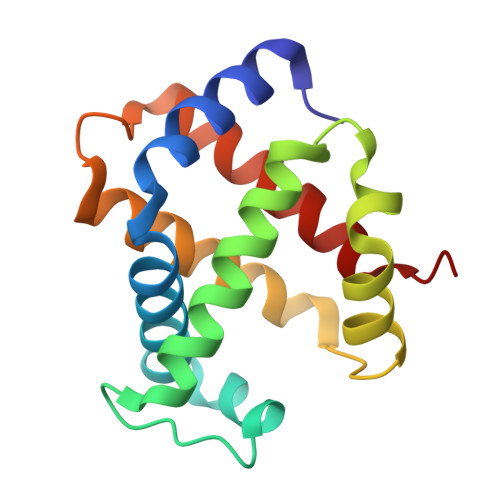
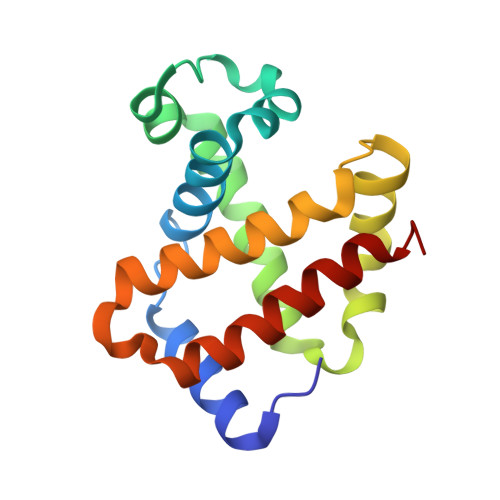
Structural and in Vitro Chracterization of Pyridyl Derivatives of Benzaldehydes : Highly Potent Antisickling Agents
Abulmalik, O., Ghatge, M.S., Nnamani, I., Eseonu, D.N., Musayev, F.N., Danso-Danquah, R., Venitz, J., Asakura, T., Abraham, D.J., Safo, M.K.To be published.