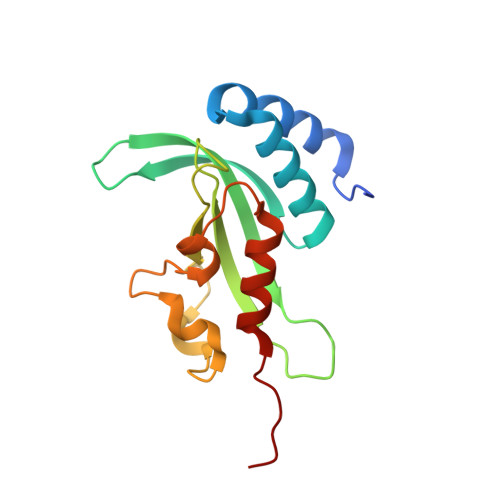
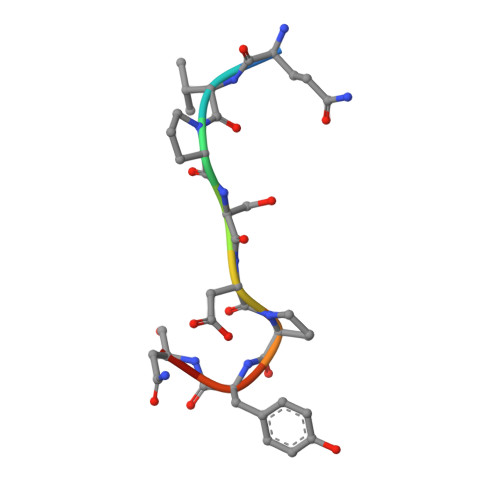
Structural basis for endosomal recruitment of ESCRT-I by ESCRT-0 in yeast.
Ren, X., Hurley, J.H.(2011) EMBO J 30: 2130-2139
- PubMed: 21505419 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.122
- Primary Citation Related Structures:
3R3Q, 3R42 - PubMed Abstract:
The ESCRT-0 and ESCRT-I complexes coordinate the clustering of ubiquitinated cargo with intralumenal budding of the endosomal membrane, two essential steps in vacuolar/lysosomal protein sorting from yeast to humans. The 1.85-Å crystal structure of interacting regions of the yeast ESCRT-0 and ESCRT-I complexes reveals that PSDP motifs of the Vps27 ESCRT-0 subunit bind to a novel electropositive N-terminal site on the UEV domain of the ESCRT-I subunit Vps23 centred on Trp16. This novel site is completely different from the C-terminal part of the human UEV domain that binds to P(S/T)AP motifs of human ESCRT-0 and HIV-1 Gag. Disruption of the novel PSDP-binding site eliminates the interaction in vitro and blocks enrichment of Vps23 in endosome-related class E compartments in yeast cells. However, this site is non-essential for sorting of the ESCRT cargo Cps1. Taken together, these results show how a conserved motif/domain pair can evolve to use strikingly different binding modes in different organisms.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD, USA.
Organizational Affiliation: