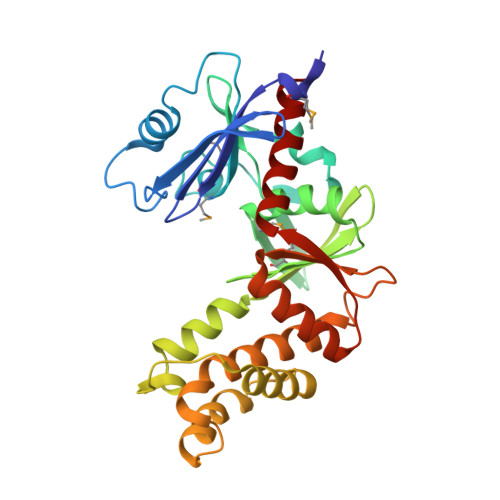
Structure of 2-oxo-3-deoxygalactonate kinase from Klebsiella pneumoniae.
Michalska, K., Cuff, M.E., Tesar, C., Feldmann, B., Joachimiak, A.(2011) Acta Crystallogr D Biol Crystallogr 67: 678-689
- PubMed: 21795809 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911021834
- Primary Citation Related Structures:
3R1X - PubMed Abstract:
In most organisms, efficient D-galactose utilization requires the highly conserved Leloir pathway that converts D-galactose to D-glucose 1-phosphate. However, in some bacterial and fungal species alternative routes of D-galactose assimilation have been identified. In the so-called De Ley-Doudoroff pathway, D-galactose is metabolized into pyruvate and D-glyceraldehyde 3-phosphate in five consecutive reactions carried out by specific enzymes. The penultimate step in this pathway involves the phosphorylation of 2-oxo-3-deoxygalactonate to 2-oxo-3-deoxygalactonate 6-phosphate catalyzed by 2-oxo-3-deoxygalactonate kinase, with ATP serving as a phosphoryl-group donor. Here, a crystal structure of 2-oxo-3-deoxygalactonate kinase from Klebsiella pneumoniae determined at 2.1 Å resolution is reported, the first structure of an enzyme from the De Ley-Doudoroff pathway. Structural comparison indicates that the enzyme belongs to the ASKHA (acetate and sugar kinases/hsc70/actin) family of phosphotransferases. The protein is composed of two α/β domains, each of which contains a core common to all family members. Additional elements introduced between conserved structural motifs define the unique features of 2-oxo-3-deoxygalactonate kinase and possibly determine the biological function of the protein.
- Midwest Center for Structural Genomics, Biosciences Division, Argonne National Laboratory, USA.
Organizational Affiliation: