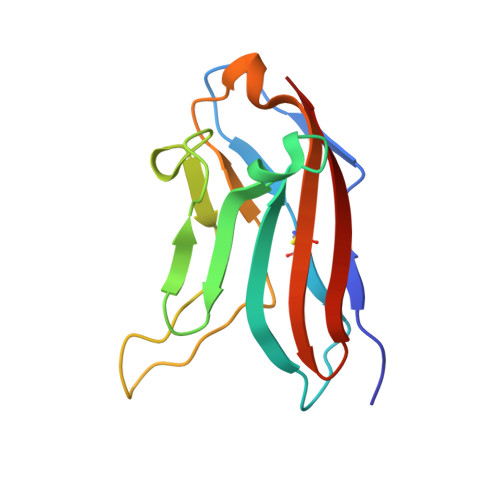
Structure of Nectin-2 reveals determinants of homophilic and heterophilic interactions that control cell-cell adhesion.
Samanta, D., Ramagopal, U.A., Rubinstein, R., Vigdorovich, V., Nathenson, S.G., Almo, S.C.(2012) Proc Natl Acad Sci U S A 109: 14836-14840
- PubMed: 22927415
- DOI: https://doi.org/10.1073/pnas.1212912109
- Primary Citation of Related Structures:
3R0N - PubMed Abstract:
Nectins are members of the Ig superfamily that mediate cell-cell adhesion through homophilic and heterophilic interactions. We have determined the crystal structure of the nectin-2 homodimer at 1.3 Å resolution. Structural analysis and complementary mutagenesis studies reveal the basis for recognition and selectivity among the nectin family members. Notably, the close proximity of charged residues at the dimer interface is a major determinant of the binding affinities associated with homophilic and heterophilic interactions within the nectin family. Our structural and biochemical data provide a mechanistic basis to explain stronger heterophilic versus weaker homophilic interactions among these family members and also offer insights into nectin-mediated transinteractions between engaging cells.
- Department of Microbiology and Immunology, Albert Einstein College of Medicine, Bronx, NY 10461, USA.
Organizational Affiliation: