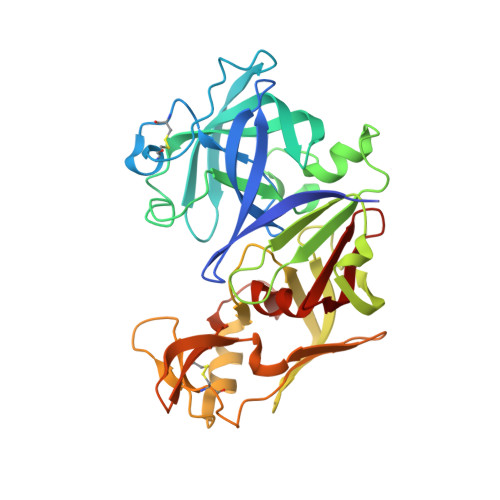
Crystal structures of the free and inhibited forms of plasmepsin I (PMI) from Plasmodium falciparum.
Bhaumik, P., Horimoto, Y., Xiao, H., Miura, T., Hidaka, K., Kiso, Y., Wlodawer, A., Yada, R.Y., Gustchina, A.(2011) J Struct Biol 175: 73-84
- PubMed: 21521654
- DOI: https://doi.org/10.1016/j.jsb.2011.04.009
- Primary Citation of Related Structures:
3QRV, 3QS1 - PubMed Abstract:
Plasmepsin I (PMI) is one of the four vacuolar pepsin-like proteases responsible for hemoglobin degradation by the malarial parasite Plasmodium falciparum, and the only one with no crystal structure reported to date. Due to substantial functional redundancy of these enzymes, lack of inhibition of even a single plasmepsin can defeat efforts in creating effective antiparasitic agents. We have now solved crystal structures of the recombinant PMI as apoenzyme and in complex with the potent peptidic inhibitor, KNI-10006, at the resolution of 2.4 and 3.1Å, respectively. The apoenzyme crystallized in the orthorhombic space group P2(1)2(1)2(1) with two molecules in the asymmetric unit and the structure has been refined to the final R-factor of 20.7%. The KNI-10006 bound enzyme crystallized in the tetragonal space group P4(3) with four molecules in the asymmetric unit and the structure has been refined to the final R-factor of 21.1%. In the PMI-KNI-10006 complex, the inhibitors were bound identically to all four enzyme molecules, with the opposite directionality of the main chain of KNI-10006 relative to the direction of the enzyme substrates. Such a mode of binding of inhibitors containing an allophenylnorstatine-dimethylthioproline insert in the P1-P1' positions, previously reported in a complex with PMIV, demonstrates the importance of satisfying the requirements for the proper positioning of the functional groups in the mechanism-based inhibitors towards the catalytic machinery of aspartic proteases, as opposed to binding driven solely by the specificity of the individual enzymes. A comparison of the structure of the PMI-KNI-10006 complex with the structures of other vacuolar plasmepsins identified the important differences between them and may help in the design of specific inhibitors targeting the individual enzymes.
- Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute, Frederick, MD 21702, USA.
Organizational Affiliation: