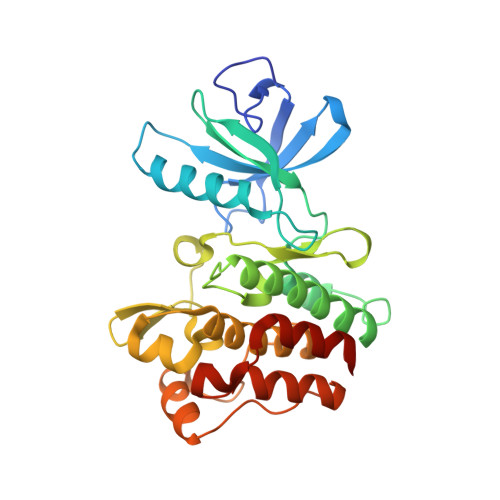
Conformational Control Inhibition of the BCR-ABL1 Tyrosine Kinase, Including the Gatekeeper T315I Mutant, by the Switch-Control Inhibitor DCC-2036.
Chan, W.W., Wise, S.C., Kaufman, M.D., Ahn, Y.M., Ensinger, C.L., Haack, T., Hood, M.M., Jones, J., Lord, J.W., Lu, W.P., Miller, D., Patt, W.C., Smith, B.D., Petillo, P.A., Rutkoski, T.J., Telikepalli, H., Vogeti, L., Yao, T., Chun, L., Clark, R., Evangelista, P., Gavrilescu, L.C., Lazarides, K., Zaleskas, V.M., Stewart, L.J., Van Etten, R.A., Flynn, D.L.(2011) Cancer Cell 19: 556-568
- PubMed: 21481795
- DOI: https://doi.org/10.1016/j.ccr.2011.03.003
- Primary Citation of Related Structures:
3QRI, 3QRJ, 3QRK - PubMed Abstract:
Acquired resistance to ABL1 tyrosine kinase inhibitors (TKIs) through ABL1 kinase domain mutations, particularly the gatekeeper mutant T315I, is a significant problem for patients with chronic myeloid leukemia (CML). Using structure-based drug design, we developed compounds that bind to residues (Arg386/Glu282) ABL1 uses to switch between inactive and active conformations. The lead "switch-control" inhibitor, DCC-2036, potently inhibits both unphosphorylated and phosphorylated ABL1 by inducing a type II inactive conformation, and retains efficacy against the majority of clinically relevant CML-resistance mutants, including T315I. DCC-2036 inhibits BCR-ABL1(T315I)-expressing cell lines, prolongs survival in mouse models of T315I mutant CML and B-lymphoblastic leukemia, and inhibits primary patient leukemia cells expressing T315I in vitro and in vivo, supporting its clinical development in TKI-resistant Ph(+) leukemia.
- Molecular Oncology Research Institute, Tufts Medical Center, Boston, MA 02111, USA.
Organizational Affiliation: