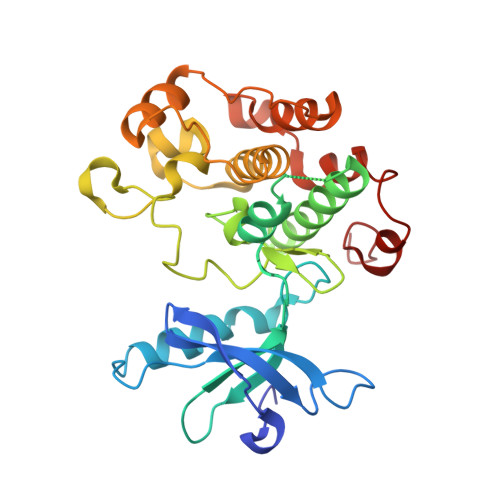
Discovery of 2,4-bis-arylamino-1,3-pyrimidines as insulin-like growth factor-1 receptor (IGF-1R) inhibitors.
Buchanan, J.L., Newcomb, J.R., Carney, D.P., Chaffee, S.C., Chai, L., Cupples, R., Epstein, L.F., Gallant, P., Gu, Y., Harmange, J.C., Hodge, K., Houk, B.E., Huang, X., Jona, J., Joseph, S., Jun, H.T., Kumar, R., Li, C., Lu, J., Menges, T., Morrison, M.J., Novak, P.M., van der Plas, S., Radinsky, R., Rose, P.E., Sawant, S., Sun, J.R., Surapaneni, S., Turci, S.M., Xu, K., Yanez, E., Zhao, H., Zhu, X.(2011) Bioorg Med Chem Lett 21: 2394-2399
- PubMed: 21414779 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.02.075
- Primary Citation Related Structures:
3QQU - PubMed Abstract:
The insulin-like growth factor-1 receptor (IGF-1R) plays an important role in the regulation of cell growth and differentiation, and in protection from apoptosis. IGF-1R has been shown to be an appealing target for the treatment of human cancer. Herein, we report the synthesis, structure-activity relationships (SAR), X-ray cocrystal structure and in vivo tumor study results for a series of 2,4-bis-arylamino-1,3-pyrimidines.
- Department of Medicinal Chemistry, Amgen Inc., 360 Binney Street, Cambridge, MA 02142, USA. john.buchanan@amgen.com
Organizational Affiliation: