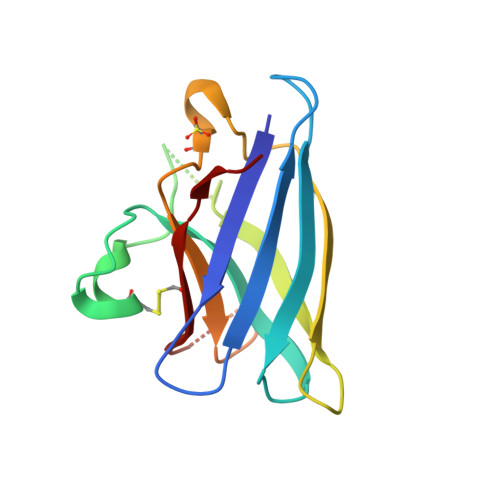
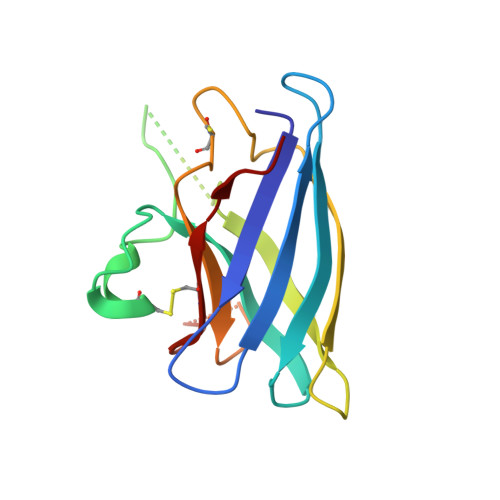
Disrupted zinc-binding sites in structures of pathogenic SOD1 variants D124V and H80R.
Seetharaman, S.V., Winkler, D.D., Taylor, A.B., Cao, X., Whitson, L.J., Doucette, P.A., Valentine, J.S., Schirf, V., Demeler, B., Carroll, M.C., Culotta, V.C., Hart, P.J.(2010) Biochemistry 49: 5714-5725
- PubMed: 20515040
- DOI: https://doi.org/10.1021/bi100314n
- Primary Citation of Related Structures:
3QQD - PubMed Abstract:
Mutations in human copper-zinc superoxide dismutase (SOD1) cause an inherited form of the fatal neurodegenerative disease amyotrophic lateral sclerosis (ALS). Here, we present structures of the pathogenic SOD1 variants D124V and H80R, both of which demonstrate compromised zinc-binding sites. The disruption of the zinc-binding sites in H80R SOD1 leads to conformational changes in loop elements, permitting non-native SOD1-SOD1 interactions that mediate the assembly of these proteins into higher-order filamentous arrays. Analytical ultracentrifugation sedimentation velocity experiments indicate that these SOD1 variants are more prone to monomerization than the wild-type enzyme. Although D124V and H80R SOD1 proteins appear to have fully functional copper-binding sites, inductively coupled plasma mass spectrometery (ICP-MS) and anomalous scattering X-ray diffraction analyses reveal that zinc (not copper) occupies the copper-binding sites in these variants. The absence of copper in these proteins, together with the results of covalent thiol modification experiments in yeast strains with and without the gene encoding the copper chaperone for SOD1 (CCS), suggests that CCS may not fully act on newly translated forms of these polypeptides. Overall, these findings lend support to the hypothesis that immature mutant SOD1 species contribute to toxicity in SOD1-linked ALS.
- Department of Biochemistry, The University of Texas Health Science Center,San Antonio, Texas 78229, USA.
Organizational Affiliation: