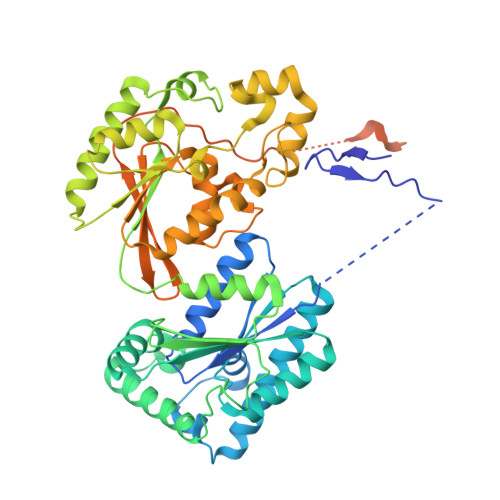
Molecular basis of the fructose-2,6-bisphosphatase reaction of PFKFB3: Transition state and the C-terminal function.
Cavalier, M.C., Kim, S.G., Neau, D., Lee, Y.H.(2012) Proteins 80: 1143-1153
- PubMed: 22275052
- DOI: https://doi.org/10.1002/prot.24015
- Primary Citation of Related Structures:
3QPU, 3QPV, 3QPW - PubMed Abstract:
The molecular basis of fructose-2,6-bisphosphatase (F-2,6-P(2)ase) of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (PFKFB) was investigated using the crystal structures of the human inducible form (PFKFB3) in a phospho-enzyme intermediate state (PFKFB3-P•F-6-P), in a transition state-analogous complex (PFKFB3•AlF(4)), and in a complex with pyrophosphate (PFKFB3•PP(i)) at resolutions of 2.45, 2.2, and 2.3 Å, respectively. Trapping the PFKFB3-P•F-6-P intermediate was achieved by flash cooling the crystal during the reaction, and the PFKFB3•AlF(4) and PFKFB3•PP(i) complexes were obtained by soaking. The PFKFB3•AlF(4) and PFKFB3•PP(i) complexes resulted in removing F-6-P from the catalytic pocket. With these structures, the structures of the Michaelis complex and the transition state were extrapolated. For both the PFKFB3-P formation and break down, the phosphoryl donor and the acceptor are located within ~5.1 Å, and the pivotal point 2-P is on the same line, suggesting an "in-line" transfer with a direct inversion of phosphate configuration. The geometry suggests that NE2 of His253 undergoes a nucleophilic attack to form a covalent N-P bond, breaking the 2O-P bond in the substrate. The resulting high reactivity of the leaving group, 2O of F-6-P, is neutralized by a proton donated by Glu322. Negative charges on the equatorial oxygen of the transient bipyramidal phosphorane formed during the transfer are stabilized by Arg252, His387, and Asn259. The C-terminal domain (residues 440-446) was rearranged in PFKFB3•PP(i), implying that this domain plays a critical role in binding of substrate to and release of product from the F-2,6-P(2) ase catalytic pocket. These findings provide a new insight into the understanding of the phosphoryl transfer reaction.
- Department of Biological Sciences, Louisiana State University, Baton Rouge, Louisiana 70803, USA.
Organizational Affiliation: