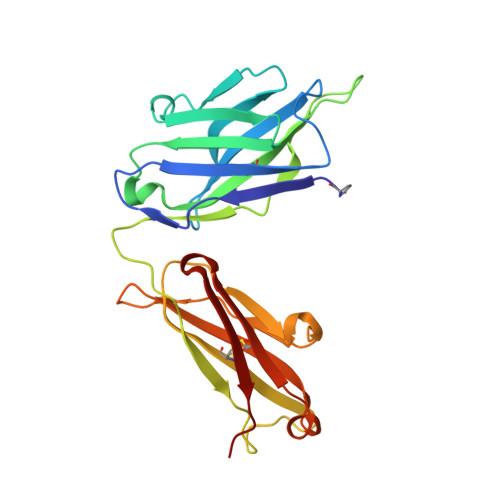
An insertion mutation that distorts antibody binding site architecture enhances function of a human antibody.
Krause, J.C., Ekiert, D.C., Tumpey, T.M., Smith, P.B., Wilson, I.A., Crowe, J.E.(2011) mBio 2: e00345-e00310
- PubMed: 21304166 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.00345-10
- Primary Citation Related Structures:
3QHF, 3QHZ - PubMed Abstract:
The structural and functional significance of somatic insertions and deletions in antibody chains is unclear. Here, we demonstrate that a naturally occurring three-amino-acid insertion within the influenza virus-specific human monoclonal antibody 2D1 heavy-chain variable region reconfigures the antibody-combining site and contributes to its high potency against the 1918 and 2009 pandemic H1N1 influenza viruses. The insertion arose through a series of events, including a somatic point mutation in a predicted hot-spot motif, introduction of a new hot-spot motif, a molecular duplication due to polymerase slippage, a deletion due to misalignment, and additional somatic point mutations. Atomic resolution structures of the wild-type antibody and a variant in which the insertion was removed revealed that the three-amino-acid insertion near the base of heavy-chain complementarity-determining region (CDR) H2 resulted in a bulge in that loop. This enlarged CDR H2 loop impinges on adjacent regions, causing distortion of the CDR H1 architecture and its displacement away from the antigen-combining site. Removal of the insertion restores the canonical structure of CDR H1 and CDR H2, but binding, neutralization activity, and in vivo activity were reduced markedly because of steric conflict of CDR H1 with the hemagglutinin antigen.
- Department of Pediatrics, Vanderbilt University Medical Center, Nashville, Tennessee, USA.
Organizational Affiliation: