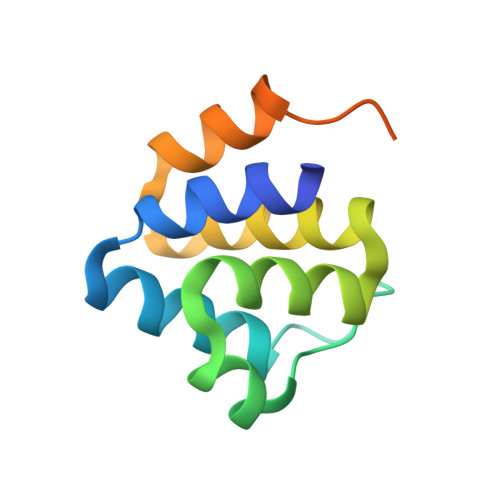
Crystal Structure of NALP3 Protein Pyrin Domain (PYD) and Its Implications in Inflammasome Assembly
Bae, J.Y., Park, H.H.(2011) J Biological Chem 286: 39528-39536
- PubMed: 21880711 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.278812
- Primary Citation Related Structures:
3QF2 - PubMed Abstract:
NALP3 inflammasome, composed of the three proteins NALP3, ASC, and Caspase-1, is a macromolecular complex responsible for the innate immune response against infection with bacterial and viral pathogens. Formation of the inflammasome can lead to the activation of inflammatory caspases, such as Caspase-1, which then activate pro-inflammatory cytokines by proteolytic cleavage. The assembly of the NALP3 inflammasome depends on the protein-interacting domain known as the death domain superfamily. NALP3 inflammasome is assembled via a pyrin domain (PYD)/PYD interaction between ASC and NALP3 and a caspase recruitment domain/caspase recruitment domain interaction between ASC and Caspase-1. As a first step toward elucidating the molecular mechanisms of inflammatory caspase activation by formation of inflammasome, we report the crystal structure of the PYD from NALP3 at 1.7-Å resolution. Although NALP3 PYD has the canonical six-helical bundle structural fold similar to other PYDs, the high resolution structure reveals the possible biologically important homodimeric interface and the dynamic properties of the fold. Comparison with other PYD structures shows both similarities and differences that may be functionally relevant. Structural and sequence analyses further implicate conserved surface residues in NALP3 PYD for ASC interaction and inflammasome assembly. The most interesting aspect of the structure was the unexpected disulfide bond between Cys-8 and Cys-108, which might be important for regulation of the activity of NALP3 by redox potential.
- Graduate School of Biochemistry, and Research Institute of Protein Sensor, Yeungnam University, Gyeongsan, South Korea.
Organizational Affiliation: