TCRs Used in Cancer Gene Therapy Cross-React with MART-1/Melan-A Tumor Antigens via Distinct Mechanisms.
Borbulevych, O.Y., Santhanagopolan, S.M., Hossain, M., Baker, B.M.(2011) J Immunol 187: 2453-2463
- PubMed: 21795600
- DOI: https://doi.org/10.4049/jimmunol.1101268
- Primary Citation of Related Structures:
3QDG, 3QDJ, 3QDM, 3QEQ, 3QEU - PubMed Abstract:
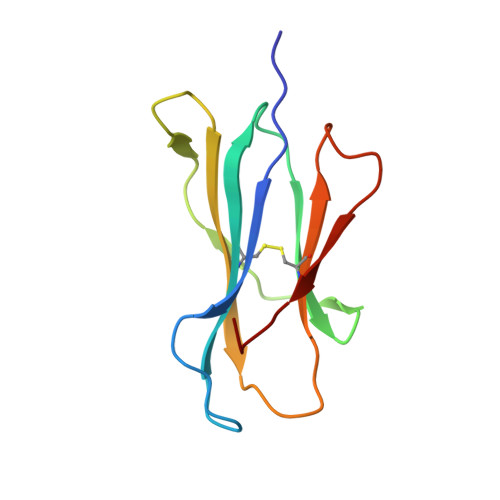
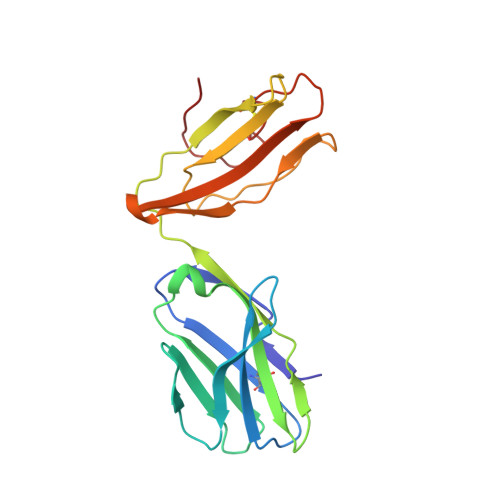
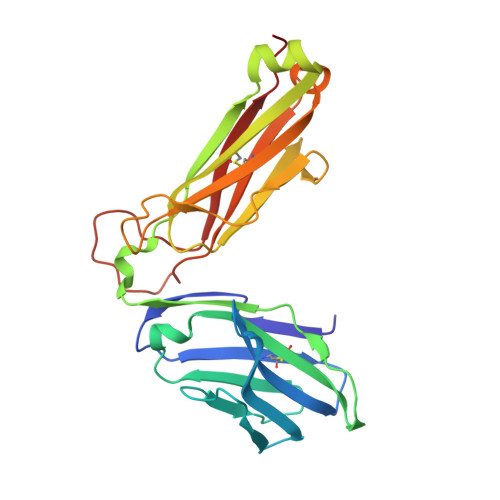
T cells engineered to express TCRs specific for tumor Ags can drive cancer regression. The first TCRs used in cancer gene therapy, DMF4 and DMF5, recognize two structurally distinct peptide epitopes of the melanoma-associated MART-1/Melan-A protein, both presented by the class I MHC protein HLA-A*0201. To help understand the mechanisms of TCR cross-reactivity and provide a foundation for the further development of immunotherapy, we determined the crystallographic structures of DMF4 and DMF5 in complex with both of the MART-1/Melan-A epitopes. The two TCRs use different mechanisms to accommodate the two ligands. Although DMF4 binds the two with a different orientation, altering its position over the peptide/MHC, DMF5 binds them both identically. The simpler mode of cross-reactivity by DMF5 is associated with higher affinity toward both ligands, consistent with the superior functional avidity of DMF5. More generally, the observation of two diverging mechanisms of cross-reactivity with the same Ags and the finding that TCR-binding orientation can be determined by peptide alone extend our understanding of the mechanisms underlying TCR cross-reactivity.
- Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, IN 46556, USA.
Organizational Affiliation: