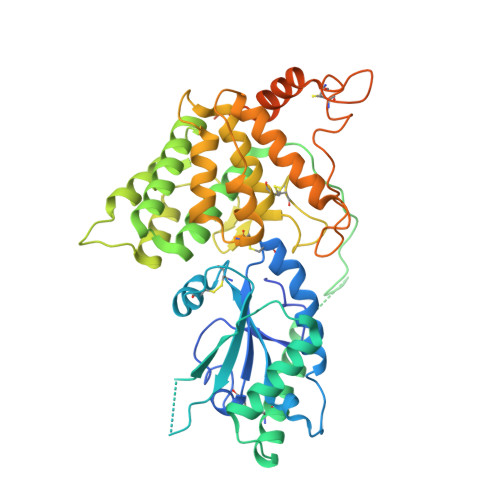
The dynamic disulphide relay of quiescin sulphydryl oxidase.
Alon, A., Grossman, I., Gat, Y., Kodali, V.K., DiMaio, F., Mehlman, T., Haran, G., Baker, D., Thorpe, C., Fass, D.(2012) Nature 488: 414-418
- PubMed: 22801504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature11267
- Primary Citation Related Structures:
3Q6O, 3QCP, 3QD9, 3T58, 3T59 - PubMed Abstract:
Protein stability, assembly, localization and regulation often depend on the formation of disulphide crosslinks between cysteine side chains. Enzymes known as sulphydryl oxidases catalyse de novo disulphide formation and initiate intra- and intermolecular dithiol/disulphide relays to deliver the disulphides to substrate proteins. Quiescin sulphydryl oxidase (QSOX) is a unique, multi-domain disulphide catalyst that is localized primarily to the Golgi apparatus and secreted fluids and has attracted attention owing to its overproduction in tumours. In addition to its physiological importance, QSOX is a mechanistically intriguing enzyme, encompassing functions typically carried out by a series of proteins in other disulphide-formation pathways. How disulphides are relayed through the multiple redox-active sites of QSOX and whether there is a functional benefit to concatenating these sites on a single polypeptide are open questions. Here we present the first crystal structure of an intact QSOX enzyme, derived from a trypanosome parasite. Notably, sequential sites in the disulphide relay were found more than 40 Å apart in this structure, too far for direct disulphide transfer. To resolve this puzzle, we trapped and crystallized an intermediate in the disulphide hand-off, which showed a 165° domain rotation relative to the original structure, bringing the two active sites within disulphide-bonding distance. The comparable structure of a mammalian QSOX enzyme, also presented here, shows further biochemical features that facilitate disulphide transfer in metazoan orthologues. Finally, we quantified the contribution of concatenation to QSOX activity, providing general lessons for the understanding of multi-domain enzymes and the design of new catalytic relays.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: